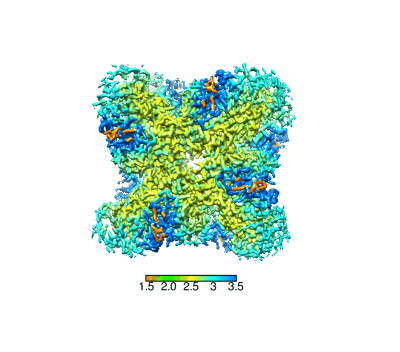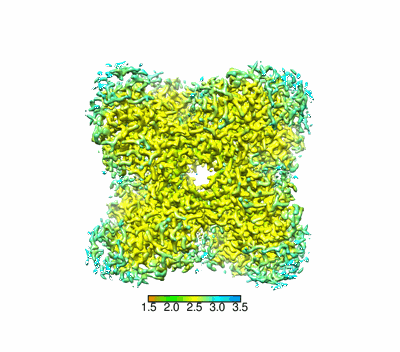Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Cryo-EM structure of GeoCas9-sgRNA-dsDNA ternary complex
Resolution: 3.08 Å
EM Method: Single-particle
Fitted PDBs: 8jtj
Q-score: 0.421
Shen PP, Liu BB, Li X, Zhang LL, Chen C-C, Guo R-T
To Be Published
- Dna (29-mer) (8 kDa, DNA from Geobacillus stearothermophilus)
- Rna (139-mer) (44 kDa, RNA from Geobacillus stearothermophilus)
- Crispr-associated endonuclease cas9 (128 kDa, Protein from Geobacillus stearothermophilus)
- Type ii crispr rna-guided endonuclease cas9 (130 kDa, Complex from Geobacillus stearothermophilus)
- Dna (5'-d(p*gp*gp*gp*cp*gp*cp*gp*ap*a)-3') (2 kDa, DNA from Geobacillus stearothermophilus)
Cryo-EM structure of GeoCas9-sgRNA binary complex
Resolution: 3.21 Å
EM Method: Single-particle
Fitted PDBs: 8jtr
Q-score: 0.401
Shen PP, Liu BB, Li X, Zhang LL, Chen C-C, Guo R-T
To Be Published
- Crispr-associated endonuclease cas9 (128 kDa, Protein from Geobacillus stearothermophilus)
- Sgrna (139-bp) (44 kDa, RNA from Geobacillus stearothermophilus)
- Type ii crispr rna-guided endonuclease cas9 (130 kDa, Complex from Geobacillus stearothermophilus)
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3
Resolution: 2.71 Å
EM Method: Single-particle
Fitted PDBs: 8ihq
Q-score: 0.632
Dai L, Niu D, Huang JW, Li X, Shen P, Li H, Xie Z, Min J, Hu Y, Yang Y, Guo RT, Chen CC
J Hazard Mater (2023) 458 pp. 131836-131836 [ Pubmed: 37331057 DOI: doi:10.1016/j.jhazmat.2023.131836 ]
- Adh3 (Complex from Stenotrophomonas acidaminiphila)
- Zinc ion (65 Da, Ligand)
- Amidohydrolase family protein (45 kDa, Protein from Stenotrophomonas acidaminiphila)
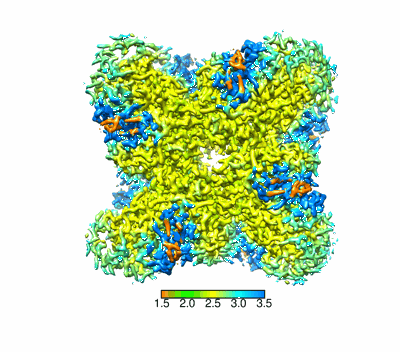
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3 in complex with Phe
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8ihr
Q-score: 0.666
Dai L, Niu D, Huang JW, Li X, Shen P, Li H, Xie Z, Min J, Hu Y, Yang Y, Guo RT, Chen CC
J Hazard Mater (2023) 458 pp. 131836-131836 [ Pubmed: 37331057 DOI: doi:10.1016/j.jhazmat.2023.131836 ]
- Adh3 (Complex from Stenotrophomonas acidaminiphila)
- Phenylalanine (165 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Amidohydrolase family protein (45 kDa, Protein from Stenotrophomonas acidaminiphila)
Cryo-EM structure of ochratoxin A-detoxifying amidohydrolase ADH3 in complex with ochratoxin A
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8ihs
Q-score: 0.645
Dai L, Niu D, Huang JW, Li X, Shen P, Li H, Xie Z, Min J, Hu Y, Yang Y, Guo RT, Chen CC
J Hazard Mater (2023) 458 pp. 131836-131836 [ Pubmed: 37331057 DOI: doi:10.1016/j.jhazmat.2023.131836 ]
- Adh3 (Complex from Stenotrophomonas acidaminiphila)
- Amidohydrolase family protein (45 kDa, Protein from Stenotrophomonas acidaminiphila)
- Zinc ion (65 Da, Ligand)
- (2~{s})-2-[[(3~{r})-5-chloranyl-3-methyl-8-oxidanyl-1-oxidanylidene-3,4-dihydroisochromen-7-yl]carbonylamino]-3-phenyl-propanoic acid (403 Da, Ligand)
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8j85
Q-score: 0.568
Dai L, Niu D, Huang JW, Li X, Shen P, Li H, Xie Z, Min J, Hu Y, Yang Y, Guo RT, Chen CC
J Hazard Mater (2023) 458 pp. 131836-131836 [ Pubmed: 37331057 DOI: doi:10.1016/j.jhazmat.2023.131836 ]
- Adh3 (Complex from Stenotrophomonas acidaminiphila)
- Zinc ion (65 Da, Ligand)
- Amidohydrolase family protein (45 kDa, Protein from Stenotrophomonas acidaminiphila)
- (2~{s})-2-[[(3~{r})-5-chloranyl-3-methyl-8-oxidanyl-1-oxidanylidene-3,4-dihydroisochromen-7-yl]carbonylamino]-3-phenyl-propanoic acid (403 Da, Ligand)