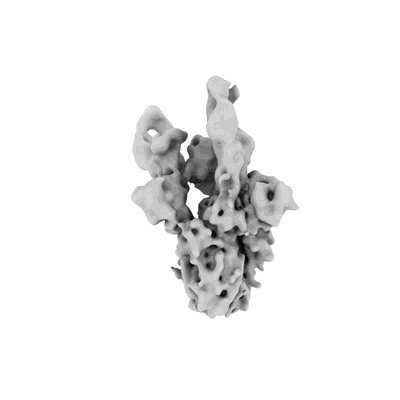Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
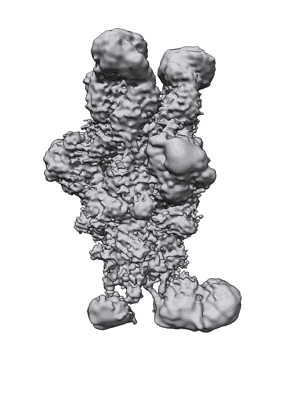
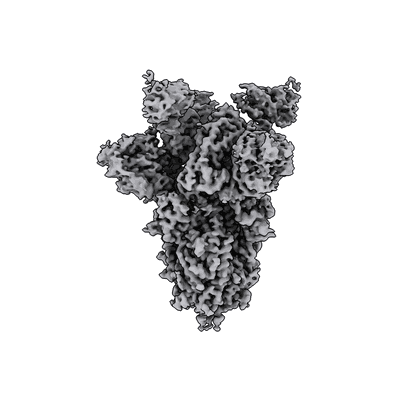
SARS-CoV-2 spike protein bound to the S2P6 and S2M11 Fab fragments
Pinto D, Sauer MM, Czudnochowski N, Low JS, Tortorici MA, Housley MP, Noack J, Walls AC, Bowen JE, Guarino B, Rosen LE, di Iulio J, Jerak J, Kaiser H, Islam S, Jaconi S, Sprugasci N, Culap K, Abdelnabi R, Foo C, Coelmont L, Bartha I, Bianchi S, Silacci-Fregni C, Bassi J, Marzi R, Vetti E, Cassotta A, Ceschi A, Ferrari P, Cippa PE, Giannini O, Ceruti S, Garzoni C, Riva A, Benigni F, Cameroni E, Piccoli L, Pizzuto MS, Smithey M, Hong D, Telenti A, Lempp FA, Neyts J, Havenar-Daughton C, Lanzavecchia A, Sallusto F, Snell G, Virgin HW, Beltramello M, Corti D, Veesler D
Science (2021) 373 pp. 1109-1116 [ Pubmed: 34344823 DOI: doi:10.1126/science.abj3321 ]
- S2m11 fab fragment (Complex from Homo sapiens)
- S2p6 fab fragment (Complex from Homo sapiens)
- S2p6 and s2m11 fab fragments bound to the sars-cov-2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
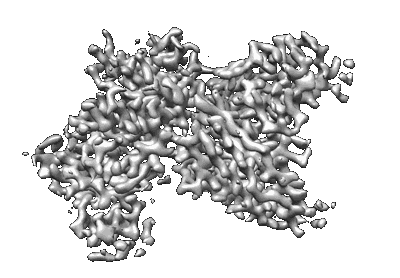
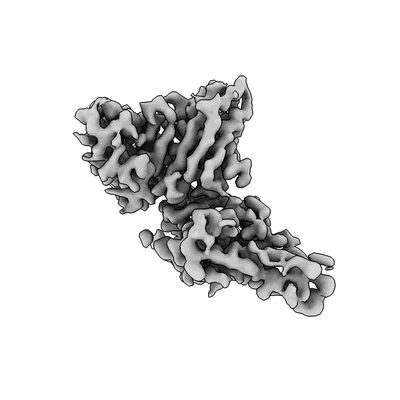
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 Global Refinement
Resolution: 2.3 Å
EM Method: Single-particle
Fitted PDBs: 7n8h
Q-score: 0.594
McCallum M, Bassi J, De Marco A, Chen A, Walls AC, Di Iulio J, Tortorici MA, Navarro MJ, Silacci-Fregni C, Saliba C, Sprouse KR, Agostini M, Pinto D, Culap K, Bianchi S, Jaconi S, Cameroni E, Bowen JE, Tilles SW, Pizzuto MS, Guastalla SB, Bona G, Pellanda AF, Garzoni C, Van Voorhis WC, Rosen LE, Snell G, Telenti A, Virgin HW, Piccoli L, Corti D, Veesler D
Science (2021) 373 pp. 648-654 [ Pubmed: 34210893 DOI: doi:10.1126/science.abi7994 ]
- S2l20 fab light chain variable region (11 kDa, Protein from Homo sapiens)
- S2l20 fab heavy chain variable region (13 kDa, Protein from Homo sapiens)
- S2m11 fab heavy chain variable region (13 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 s (b.1.429 / epsilon variant) bound to s2m11 and s2l20 fabs (Complex from Homo sapiens)
- S2m11 fab light chain variable region (11 kDa, Protein from Homo sapiens)
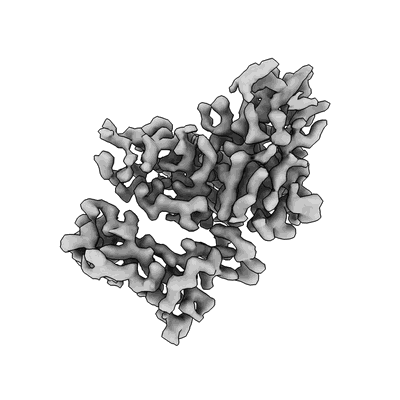
SARS-CoV-2 S (B.1.429 / epsilon variant) + S2M11 + S2L20 (Local Refinement of the NTD/S2L20)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7n8i
Q-score: 0.582
McCallum M, Bassi J, De Marco A, Chen A, Walls AC, Di Iulio J, Tortorici MA, Navarro MJ, Silacci-Fregni C, Saliba C, Sprouse KR, Agostini M, Pinto D, Culap K, Bianchi S, Jaconi S, Cameroni E, Bowen JE, Tilles SW, Pizzuto MS, Guastalla SB, Bona G, Pellanda AF, Garzoni C, Van Voorhis WC, Rosen LE, Snell G, Telenti A, Virgin HW, Piccoli L, Corti D, Veesler D
Science (2021) 373 pp. 648-654 [ Pubmed: 34210893 DOI: doi:10.1126/science.abi7994 ]
- S2l20 fab light chain variable region (11 kDa, Protein from Homo sapiens)
- S2l20 fab heavy chain variable region (13 kDa, Protein from Homo sapiens)
- Sars-cov-2 s (b.1.429 / epsilon variant) bound to s2l20 fab (Complex from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7jv2
Q-score: 0.445
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
- S2h13 neutralizing antibody fab fragment (Complex from Homo sapiens)
- S2h13 fab light chain (11 kDa, Protein from Homo sapiens)
- Sars-cov-2 spike in complex with the s2h13 neutralizing antibody fab fragment (Complex)
- Spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2h13 fab heavy chain (13 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
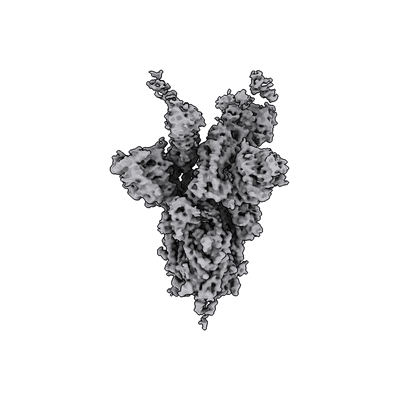
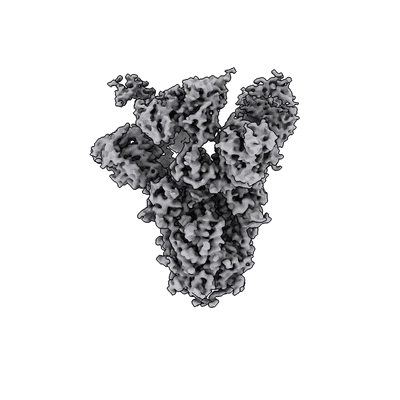
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (one RBD open)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7jv4
Q-score: 0.397
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
- S2h13 neutralizing antibody fab fragment (Complex from Homo sapiens)
- S2h13 fab heavy chain (13 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 spike in complex with the s2h13 neutralizing antibody fab fragment (Complex)
- Spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2h13 fab light chain (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (closed conformation)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7jv6
Q-score: 0.433
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
- S2h13 neutralizing antibody fab fragment (Complex from Homo sapiens)
- S2h13 fab heavy chain (13 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 spike in complex with the s2h13 neutralizing antibody fab fragment (Complex)
- Spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2h13 fab light chain (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7jva
Q-score: 0.452
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
- S2a4 fab light chain (11 kDa, Protein from Homo sapiens)
- Sars-cov-2 spike in complex with the s2a4 neutralizing antibody fab fragment (Complex)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2a4 neutralizing antibody fab fragment (Complex from Homo sapiens)
- S2a4 fab heavy chain (13 kDa, Protein from Homo sapiens)
SARS-CoV-2 spike in complex with the S2A4 neutralizing antibody Fab fragment
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7jvc
Q-score: 0.391
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
- Sars-cov-2 spike in complex with the s2a4 neutralizing antibody fab fragment (Complex)
- S2a4 fab heavy chain (24 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- S2a4 neutralizing antibody fab fragment (Complex from Homo sapiens)
- S2a4 fab light chain (23 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]
Piccoli L, Park YJ, Tortorici MA, Czudnochowski N, Walls AC, Beltramello M, Silacci-Fregni C, Pinto D, Rosen LE, Bowen JE, Acton OJ, Jaconi S, Guarino B, Minola A, Zatta F, Sprugasci N, Bassi J, Peter A, De Marco A, Nix JC, Mele F, Jovic S, Rodriguez BF, Gupta SV, Jin F, Piumatti G, Lo Presti G, Pellanda AF, Biggiogero M, Tarkowski M, Pizzuto MS, Cameroni E, Havenar-Daughton C, Smithey M, Hong D, Lepori V, Albanese E, Ceschi A, Bernasconi E, Elzi L, Ferrari P, Garzoni C, Riva A, Snell G, Sallusto F, Fink K, Virgin HW, Lanzavecchia A, Corti D, Veesler D
Cell (2020) 183 pp. 1024-1042.e21 [ DOI: doi:10.1016/j.cell.2020.09.037 Pubmed: 32991844 ]