Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
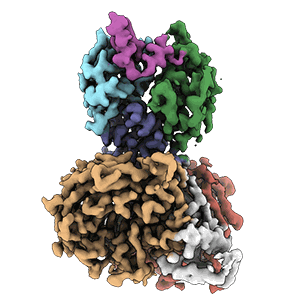
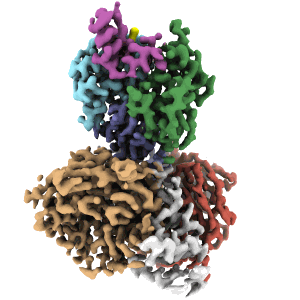
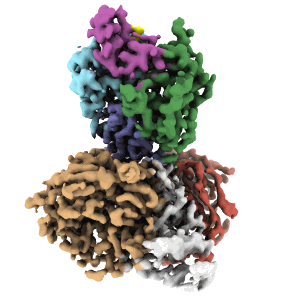
Cryo-EM structure of DDB1deltaB-DDA1-DCAF5
Resolution: 2.63 Å
EM Method: Single-particle
Fitted PDBs: 8tl6
Q-score: 0.598
Radko-Juettner S, Yue H, Myers JA, Carter RD, Robertson AN, Mittal P, Zhu Z, Hansen BS, Donovan KA, Hunkeler M, Rosikiewicz W, Wu Z, McReynolds MG, Roy Burman SS, Schmoker AM, Mageed N, Brown SA, Mobley RJ, Partridge JF, Stewart EA, Pruett-Miller SM, Nabet B, Peng J, Gray NS, Fischer ES, Roberts CWM
Nature (2024) 628 pp. 442-449 [ DOI: doi:10.1038/s41586-024-07250-1 Pubmed: 38538798 ]
- Complex of ddb1deltab-dda1-dcaf5 (220 kDa, Complex from Homo sapiens)
- Det1- and ddb1-associated protein 1 (14 kDa, Protein from Homo sapiens)
- Dna damage-binding protein 1 (96 kDa, Protein from Homo sapiens)
- Ddb1- and cul4-associated factor 5 (108 kDa, Protein from Homo sapiens)
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8tnp
Q-score: 0.507
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
- Dna damage-binding protein 1 (96 kDa, Protein from Homo sapiens)
- Ternary complex of ddb1db:crbn:pomalidomide with sd40 (201 kDa, Complex)
- S-pomalidomide (273 Da, Ligand)
- Protein cereblon (55 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Maltose/maltodextrin-binding periplasmic protein,sd40 (50 kDa, Protein from Homo sapiens)
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Resolution: 2.41 Å
EM Method: Single-particle
Fitted PDBs: 8tnq
Q-score: 0.591
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
- Dna damage-binding protein 1 (96 kDa, Protein from Homo sapiens)
- Ternary complex of ddb1db:crbn:pt-179 with sd40 (201 kDa, Complex)
- Protein cereblon (55 kDa, Protein from Homo sapiens)
- Maltose/maltodextrin-binding periplasmic protein,sd40 (50 kDa, Protein from Homo sapiens)
- 2-[(3s)-2,6-dioxopiperidin-3-yl]-5-(morpholin-4-yl)-1h-isoindole-1,3(2h)-dione (343 Da, Ligand)
- Zinc ion (65 Da, Ligand)
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8tnr
Q-score: 0.561
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
- Dna damage-binding protein 1 (96 kDa, Protein from Homo sapiens)
- Ternary complex of ddb1db:crbn:pt-179 with sd40 (201 kDa, Complex)
- Protein cereblon (55 kDa, Protein from Homo sapiens)
- Maltose/maltodextrin-binding periplasmic protein,sd40 (50 kDa, Protein from Homo sapiens)
- 2-[(3s)-2,6-dioxopiperidin-3-yl]-5-(morpholin-4-yl)-1h-isoindole-1,3(2h)-dione (343 Da, Ligand)
- Zinc ion (65 Da, Ligand)
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2
Mercer JAM, DeCarlo SJ, Roy Burman SS, Sreekanth V, Nelson AT, Hunkeler M, Chen PJ, Donovan KA, Kokkonda P, Tiwari PK, Shoba VM, Deb A, Choudhary A, Fischer ES, Liu DR
Science (2024) 383 pp. eadk4422-eadk4422 [ Pubmed: 38484051 DOI: doi:10.1126/science.adk4422 ]
Cryo-EM structure of full-length human UBR5 (homotetramer)
Resolution: 3.87 Å
EM Method: Single-particle
Fitted PDBs: 8p83
Q-score: 0.2
Tsai JM, Aguirre JD, Li YD, Brown J, Focht V, Kater L, Kempf G, Sandoval B, Schmitt S, Rutter JC, Galli P, Sandate CR, Cutler JA, Zou C, Donovan KA, Lumpkin RJ, Cavadini S, Park PMC, Sievers Q, Hatton C, Ener E, Regalado BD, Sperling MT, Slabicki M, Kim J, Zon R, Zhang Z, Miller PG, Belizaire R, Sperling AS, Fischer ES, Irizarry R, Armstrong SA, Thoma NH, Ebert BL
Mol Cell (2023) 83 pp. 2753- [ DOI: doi:10.1016/j.molcel.2023.06.028 Pubmed: 37478846 ]
- Tetramer of ubr5 (Complex from Homo sapiens)
- E3 ubiquitin-protein ligase ubr5 (313 kDa, Protein from Homo sapiens)
Negative stain map of UBR5 (dimer) in complex with RARA/RXRA
Tsai JM, Aguirre JD, Li YD, Brown J, Focht V, Kater L, Kempf G, Sandoval B, Schmitt S, Rutter JC, Galli P, Sandate CR, Cutler JA, Zou C, Donovan KA, Lumpkin RJ, Cavadini S, Park PMC, Sievers Q, Hatton C, Ener E, Regalado BD, Sperling MT, Slabicki M, Kim J, Zon R, Zhang Z, Miller PG, Belizaire R, Sperling AS, Fischer ES, Irizarry R, Armstrong SA, Thoma NH, Ebert BL
Mol Cell (2023) 83 pp. 2753-2767.e10 [ DOI: doi:10.1016/j.molcel.2023.06.028 Pubmed: 37478846 ]
- Rara (Protein from Homo sapiens)
- Ubr5 (Protein from Homo sapiens)
- Rxra (Protein from Homo sapiens)
- Ubr5 in complex with rara/rxra (Complex from Homo sapiens)
Cryo-EM structure of DDB1deltaB-DDA1-DCAF16-BRD4(BD2)-MMH2
Resolution: 2.2 Å
EM Method: Single-particle
Fitted PDBs: 8g46
Q-score: 0.643
Li YD, Ma MW, Hassan MM, Hunkeler M, Teng M, Puvar K, Lumpkin R, Sandoval B, Jin CY, Ficarro SB, Wang MY, Xu S, Groendyke BJ, Sigua LH, Tavares I, Zou C, Tsai JM, Park PMC, Yoon H, Majewski FC, Marto JA, Qi J, Nowak RP, Donovan KA, Slabicki M, Gray NS, Fischer ES, Ebert BL
bioRxiv (2023) [ Pubmed: 36824856 DOI: doi:10.1101/2023.02.14.528208 ]
- Bromodomain-containing protein 4 (14 kDa, Protein from Homo sapiens)
- Dna damage-binding protein 1 (96 kDa, Protein from Homo sapiens)
- Det1- and ddb1-associated protein 1 (12 kDa, Protein from Homo sapiens)
- Tert-butyl [(6s,10p)-4-{4-[(ethanesulfonyl)amino]phenyl}-2,3,9-trimethyl-6h-thieno[3,2-f][1,2,4]triazolo[4,3-a][1,4]diazepin-6-yl]acetate (529 Da, Ligand)
- Water (18 Da, Ligand)
- Ternary complex of cullin ring e3 ubiquitin ligase substrate receptor arm (ddb1deltab-dda1-dcaf16) in compound-induced complex with bromodomain 2 of brd4 (148 kDa, Complex from Homo sapiens)
- Ddb1- and cul4-associated factor 16 (24 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)