Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8v4y
Q-score: 0.53
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h3.2 (15 kDa, Protein from Xenopus laevis)
- Beryllium trifluoride ion (66 Da, Ligand)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (64 kDa, DNA from synthetic construct)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Magnesium ion (24 Da, Ligand)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Histone h2a type 1 (13 kDa, Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (122 kDa, Protein from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (63 kDa, DNA from synthetic construct)
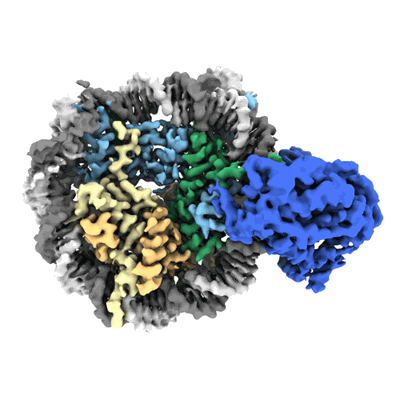
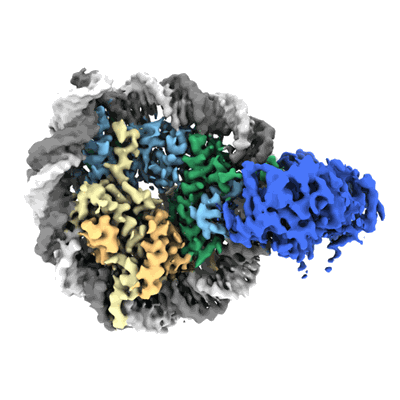
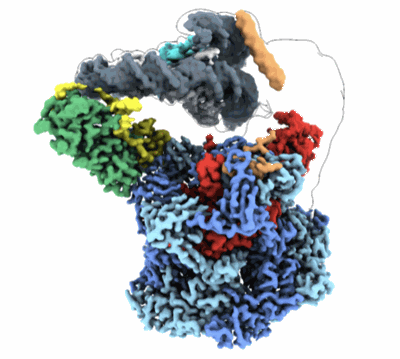
Cryo-EM structure of SNF2h-nucleosome complex (consensus structure)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h4 (Protein from Xenopus laevis)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna reverse (DNA from synthetic construct)
- Nucleosome with xenopus laevis histones (Complex from Xenopus laevis)
- Snf2h (Complex from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (100 kDa, Complex from Homo sapiens)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna forward (DNA from synthetic construct)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Histone h2b 1.1 (Protein from Xenopus laevis)
- Histone h3.2 (Protein from Xenopus laevis)
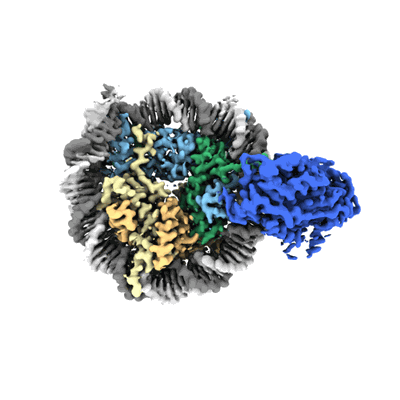
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h4 (Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna forward strand (DNA from synthetic construct)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna reverse strand (DNA from synthetic construct)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Histone h2b 1.1 (Protein from Xenopus laevis)
- Histone h3.2 (Protein from Xenopus laevis)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
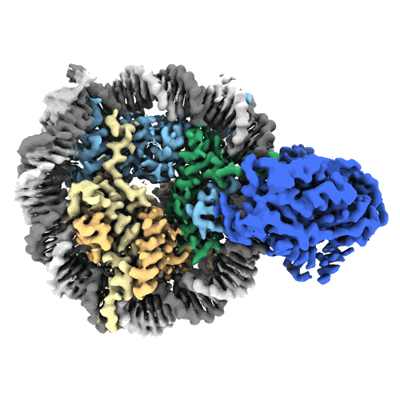
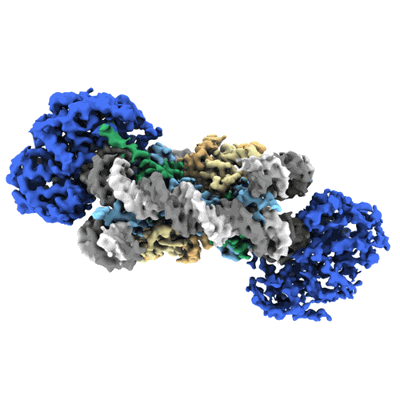
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8v6v
Q-score: 0.477
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h3.2 (15 kDa, Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (64 kDa, DNA from synthetic construct)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Magnesium ion (24 Da, Ligand)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Histone h2a type 1 (13 kDa, Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (122 kDa, Protein from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (63 kDa, DNA from synthetic construct)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8v7l
Q-score: 0.456
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h3.2 (15 kDa, Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Magnesium ion (24 Da, Ligand)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Histone h2a type 1 (13 kDa, Protein from Xenopus laevis)
- Widom 601 dna (147-mer) plus 60 base pairs flanking dna (reverse strand) (64 kDa, DNA from synthetic construct)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (122 kDa, Protein from Homo sapiens)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (63 kDa, DNA from synthetic construct)
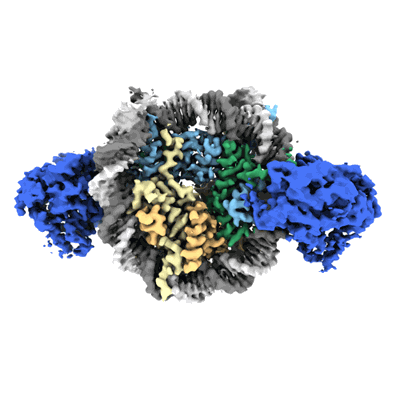
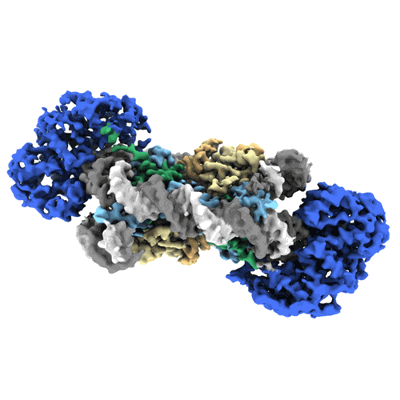
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h4 (Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (DNA from synthetic construct)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (DNA from synthetic construct)
- Histone h2b 1.1 (Protein from Xenopus laevis)
- Histone h3.2 (Protein from Xenopus laevis)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)
Chio US, Palovcak E, Smith AAA, Autzen H, Munoz EN, Yu Z, Wang F, Agard DA, Armache JP, Narlikar GJ, Cheng Y
Nat Commun (2024) 15 pp. 2225-2225 [ Pubmed: 38472177 DOI: doi:10.1038/s41467-024-46178-y ]
- Histone h4 (Protein from Xenopus laevis)
- Nucleosome with xenopus laevis histones (200 kDa, Complex)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (reverse strand) (DNA from synthetic construct)
- Snf2h (100 kDa, Complex from Homo sapiens)
- Histone h2a type 1 (Protein from Xenopus laevis)
- Swi/snf-related matrix-associated actin-dependent regulator of chromatin subfamily a member 5 (Protein from Homo sapiens)
- Widom 601 dna (147-mer) with 60 base pairs flanking dna (forward strand) (DNA from synthetic construct)
- Histone h2b 1.1 (Protein from Xenopus laevis)
- Histone h3.2 (Protein from Xenopus laevis)
- Atp-dependent chromatin remodeler snf2h in complex with nucleosome (300 kDa, Complex)
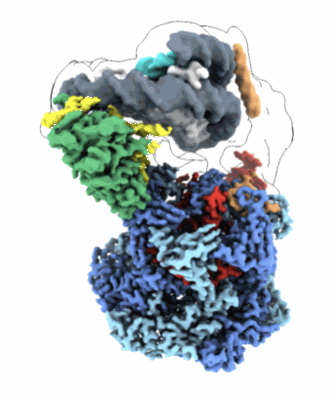
Cryo-EM structure of the human Sirtuin 6-nucleosome complex
Chio US, Rechiche O, Bryll AR, Zhu J, Leith EM, Feldman JL, Peterson CL, Tan S, Armache JP
Sci Adv (2023) 9 [ DOI: 10.1126/sciadv.adf7586 Pubmed: 37058572 ]
Image sets:
Class1 of the INO80-Hexasome complex
Resolution: 3.04 Å
EM Method: Single-particle
Fitted PDBs: 8ets
Q-score: 0.473
Wu H, Munoz EN, Hsieh LJ, Chio US, Gourdet MA, Narlikar GJ, Cheng Y
Science (2023) 381 pp. 319-324 [ Pubmed: 37384669 DOI: doi:10.1126/science.adf4197 ]
- Ino eighty subunit 2 (3 kDa, Protein from baker's yeast)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ruvb-like protein 1 (48 kDa, Protein from baker's yeast)
- Actin-related protein 5 (86 kDa, Protein from baker's yeast)
- Chromatin-remodeling atpase ino80 (54 kDa, Protein from baker's yeast)
- Chromatin-remodeling complex subunit ies6 (15 kDa, Protein from baker's yeast)
- Ino80-hex (Complex from Saccharomyces cerevisiae)
- Ruvb-like protein 2 (50 kDa, Protein from baker's yeast)
Class2 of the INO80-Hexasome complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8etu
Q-score: 0.512
Wu H, Munoz EN, Hsieh LJ, Chio US, Gourdet MA, Narlikar GJ, Cheng Y
Science (2023) 381 pp. 319-324 [ Pubmed: 37384669 DOI: doi:10.1126/science.adf4197 ]
- Ino eighty subunit 2 (3 kDa, Protein from baker's yeast)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ruvb-like protein 1 (48 kDa, Protein from baker's yeast)
- Actin-related protein 5 (86 kDa, Protein from baker's yeast)
- Chromatin-remodeling atpase ino80 (54 kDa, Protein from baker's yeast)
- Chromatin-remodeling complex subunit ies6 (15 kDa, Protein from baker's yeast)
- Ino80-hex (Complex from Saccharomyces cerevisiae)
- Ruvb-like protein 2 (50 kDa, Protein from baker's yeast)