Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
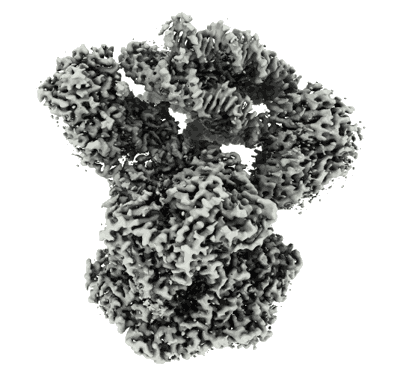
Cryo-EM structure of the PP2A:B55-FAM122A complex, B55 body
Resolution: 2.55 Å
EM Method: Single-particle
Fitted PDBs: 8twe
Q-score: 0.592
Padi SKR, Vos MR, Godek RJ, Fuller JR, Kruse T, Hein JB, Nilsson J, Kelker MS, Page R, Peti W
Nature (2024) 625 pp. 195-203 [ DOI: doi:10.1038/s41586-023-06870-3 Pubmed: 38123684 ]
- Quadruple complex of pp2a:b55 (pp2aa:pp2ac:b55) bound to fam122a (Complex)
- Serine/threonine-protein phosphatase 2a 55 kda regulatory subunit b alpha isoform (52 kDa, Protein from Homo sapiens)
- Ppp2r1a-ppp2r2a-interacting phosphatase regulator 1 (10 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a 65 kda regulatory subunit a alpha isoform (64 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a catalytic subunit alpha isoform (35 kDa, Protein from Homo sapiens)
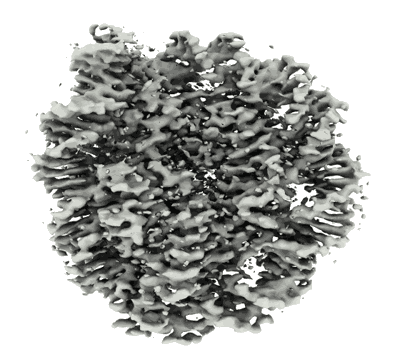
Cryo-EM structure of the PP2A:B55-FAM122A complex, PP2Ac body
Resolution: 2.69 Å
EM Method: Single-particle
Fitted PDBs: 8twi
Q-score: 0.499
Padi SKR, Vos MR, Godek RJ, Fuller JR, Kruse T, Hein JB, Nilsson J, Kelker MS, Page R, Peti W
Nature (2024) 625 pp. 195-203 [ DOI: doi:10.1038/s41586-023-06870-3 Pubmed: 38123684 ]
- Quadruple complex of pp2a:b55 (pp2aa:pp2ac:b55) bound to fam122a (Complex)
- Fe (iii) ion (55 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Ppp2r1a-ppp2r2a-interacting phosphatase regulator 1 (10 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a 65 kda regulatory subunit a alpha isoform (64 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a catalytic subunit alpha isoform (35 kDa, Protein from Homo sapiens)
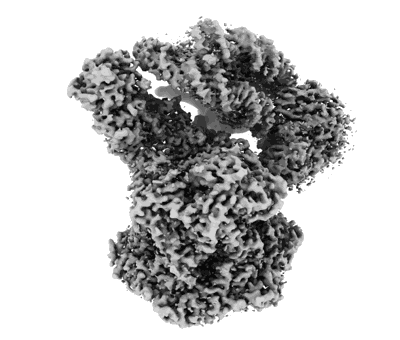
Cryo-EM structure of the PP2A:B55-FAM122A complex
Resolution: 2.79961 Å
EM Method: Single-particle
Fitted PDBs: 8so0
Q-score: 0.561
Padi SKR, Vos MR, Godek RJ, Fuller JR, Kruse T, Hein JB, Nilsson J, Kelker MS, Page R, Peti W
Nature (2024) 625 pp. 195-203 [ DOI: doi:10.1038/s41586-023-06870-3 Pubmed: 38123684 ]
- Quadruple complex of pp2a:b55 (pp2aa:pp2ac:b55) bound to fam122a (Complex)
- Fe (iii) ion (55 Da, Ligand)
- Serine/threonine-protein phosphatase 2a 55 kda regulatory subunit b alpha isoform (52 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Ppp2r1a-ppp2r2a-interacting phosphatase regulator 1 (10 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a 65 kda regulatory subunit a alpha isoform (64 kDa, Protein from Homo sapiens)
- Serine/threonine-protein phosphatase 2a catalytic subunit alpha isoform (35 kDa, Protein from Homo sapiens)
Cryo-EM Structure of Human Serine Hydroxymethyltransferase, isoform 2 (SHMT2)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8qi7
Q-score: 0.598
Rutkiewicz M, Tran LH, Ruszkowski M
To Be Published
- Water (18 Da, Ligand)
- Serine hydroxymethyltransferase, mitochondrial (52 kDa, Protein from Homo sapiens)
- Shmt2 in the form of plp internal aldimine (Complex from Homo sapiens)
CryoEM Structure INO80core Hexasome complex composite map state1
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8oo7
Q-score: 0.556
Zhang M, Jungblut A, Kunert F, Hauptmann L, Hoffmann T, Kolesnikova O, Metzner F, Moldt M, Weis F, DiMaio F, Hopfner KP, Eustermann S
Science (2023) 381 pp. 313-319 [ Pubmed: 37384673 DOI: doi:10.1126/science.adf6287 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ino80 core module in complex with hexasome (861 kDa, Complex)
- Histone h2a (14 kDa, Protein from Homo sapiens)
- Histone h2b (13 kDa, Protein from Homo sapiens)
- Dna strand 1 (69 kDa, DNA from synthetic construct)
- Dna (Complex from Synthetic construct)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Dna strand 2 (70 kDa, DNA from synthetic construct)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Actin-related protein 5 (87 kDa, Protein from Thermochaetoides thermophila)
- Ruvb-like protein 1 (50 kDa, Protein from Thermochaetoides thermophila)
- Chromatin remodeler ino80 (Complex from Thermochaetoides thermophila)
- Chromatin-remodeling complex subunit ies6 (23 kDa, Protein from Thermochaetoides thermophila)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Chromatin-remodeling atpase ino80 (130 kDa, Protein from Thermochaetoides thermophila)
- Ino eighty subunit 2 (53 kDa, Protein from Thermochaetoides thermophila)
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Ruvb-like protein 2 (53 kDa, Protein from Thermochaetoides thermophila)
- Magnesium ion (24 Da, Ligand)
- Histones (Complex from Homo sapiens)
CryoEM Structure INO80core Hexasome complex Hexasome refinement state1
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 8ooa
Q-score: 0.525
Zhang M, Jungblut A, Kunert F, Hauptmann L, Hoffmann T, Kolesnikova O, Metzner F, Moldt M, Weis F, DiMaio F, Hopfner KP, Eustermann S
Science (2023) 381 pp. 313-319 [ Pubmed: 37384673 DOI: doi:10.1126/science.adf6287 ]
- Histone h2a (14 kDa, Protein from Homo sapiens)
- Dna strand 1 (69 kDa, DNA from synthetic construct)
- Histone h2b (13 kDa, Protein from Homo sapiens)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Ino80 core module in complex with hexasome (861 kDa, Complex from Thermochaetoides thermophila)
- Dna strand 2 (70 kDa, DNA from synthetic construct)
INO80 core bound to hexasome composite map of state 2
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8oop
Q-score: 0.554
Zhang M, Jungblut A, Kunert F, Hauptmann L, Hoffmann T, Kolesnikova O, Metzner F, Moldt M, Weis F, DiMaio F, Hopfner KP, Eustermann S
Science (2023) 381 pp. 313-319 [ Pubmed: 37384673 DOI: doi:10.1126/science.adf6287 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ino80 core module in complex with hexasome (861 kDa, Complex)
- Histone h2a (14 kDa, Protein from Homo sapiens)
- Dna strand 1 (69 kDa, DNA from synthetic construct)
- Histone h2b (13 kDa, Protein from Homo sapiens)
- Dna (Complex from Synthetic construct)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Dna strand 2 (70 kDa, DNA from synthetic construct)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Actin-related protein 5 (87 kDa, Protein from Thermochaetoides thermophila)
- Ruvb-like protein 1 (50 kDa, Protein from Thermochaetoides thermophila)
- Chromatin remodeler ino80 (Complex from Thermochaetoides thermophila)
- Chromatin-remodeling complex subunit ies6 (23 kDa, Protein from Thermochaetoides thermophila)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Chromatin-remodeling atpase ino80 (130 kDa, Protein from Thermochaetoides thermophila)
- Ino eighty subunit 2 (53 kDa, Protein from Thermochaetoides thermophila)
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Ruvb-like protein 2 (53 kDa, Protein from Thermochaetoides thermophila)
- Magnesium ion (24 Da, Ligand)
- Histones (Complex from Homo sapiens)
CryoEM Structure INO80core Hexasome complex ATPase-hexasome refinement state 2
Resolution: 3.29 Å
EM Method: Single-particle
Fitted PDBs: 8oos
Q-score: 0.494
Zhang M, Jungblut A, Kunert F, Hauptmann L, Hoffmann T, Kolesnikova O, Metzner F, Moldt M, Weis F, DiMaio F, Hopfner KP, Eustermann S
Science (2023) 381 pp. 313-319 [ Pubmed: 37384673 DOI: doi:10.1126/science.adf6287 ]
- Histone h2a (14 kDa, Protein from Homo sapiens)
- Dna strand 1 (69 kDa, DNA from synthetic construct)
- Histone h2b (13 kDa, Protein from Homo sapiens)
- Ino80 (Complex from Thermochaetoides thermophila)
- Chromatin-remodeling atpase ino80 (130 kDa, Protein from Thermochaetoides thermophila)
- Dna (Complex from Synthetic construct)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Dna strand 2 (70 kDa, DNA from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Histones (Complex from Homo sapiens)
- Ino80 core module in complex with hexasome (861 kDa, Complex)
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8g8f
Q-score: 0.573
O'Neill AG, Burrell AL, Zech M, Elpeleg O, Harel T, Edvardson S, Mor-Shaked H, Rippert AL, Nomakuchi T, Izumi K, Kollman JM
J Biol Chem (2023) 299 pp. 105012-105012 [ Pubmed: 37414152 DOI: doi:10.1016/j.jbc.2023.105012 ]
- Inosine-5'-monophosphate dehydrogenase 2 (56 kDa, Protein from Homo sapiens)
- Filament of inosine-5'-monophosphate dehydrogenase 2 - l245p mutant, bound to atp, imp, and nad+ (Complex from Homo sapiens)
- Nicotinamide-adenine-dinucleotide (663 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Inosinic acid (348 Da, Ligand)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8g9b
Q-score: 0.529
O'Neill AG, Burrell AL, Zech M, Elpeleg O, Harel T, Edvardson S, Mor-Shaked H, Rippert AL, Nomakuchi T, Izumi K, Kollman JM
J Biol Chem (2023) 299 pp. 105012-105012 [ Pubmed: 37414152 DOI: doi:10.1016/j.jbc.2023.105012 ]
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Inosine-5'-monophosphate dehydrogenase 2 (56 kDa, Protein from Homo sapiens)
- Nicotinamide-adenine-dinucleotide (663 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Inosinic acid (348 Da, Ligand)
- Filament of inosine-5'-monophosphate dehydrogenase 2 - l245p mutant, bound to gtp, atp, imp, and nad+ (Complex from Homo sapiens)