Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
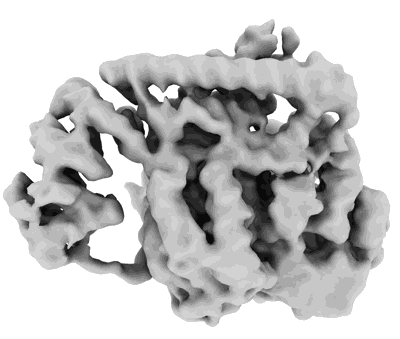
Porcine epidemic diarrhea virus core polymerase complex
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8g6r
Q-score: 0.435
Anderson TK, Hoferle PJ, Lee KW, Coon JJ, Kirchdoerfer RN
bioRxiv (2023) [ DOI: doi:10.1101/2023.03.15.532841 Pubmed: 36993498 ]
- Core replication complex of porcine epidemic diarrhea virus composed of nsp12, nsp7, nsp8, and a short rna substrate (185 kDa, Complex from Porcine epidemic diarrhea virus)
- Nsp7 (6 kDa, Protein from Porcine epidemic diarrhea virus)
- Nsp8 (12 kDa, Protein from Porcine epidemic diarrhea virus)
- Zinc ion (65 Da, Ligand)
- Rna (5'-r(p*ap*ap*gp*ap*ap*gp*cp*up*ap*up*up*ap*ap*ap*ap*up*cp*ap*cp*a)-3') (6 kDa, RNA from synthetic construct)
- Rna (5'-r(p*gp*gp*up*up*gp*up*gp*ap*up*up*up*up*ap*ap*up*ap*gp*cp*up*u)-3') (6 kDa, RNA from synthetic construct)
- Nsp12 (104 kDa, Protein from Porcine epidemic diarrhea virus)
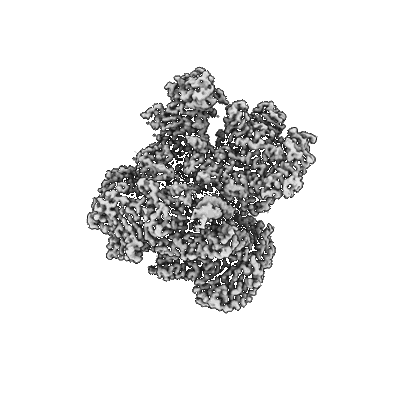
Cryo-EM structure of POLRMT mutant.
Resolution: 3.24 Å
EM Method: Single-particle
Fitted PDBs: 7zc4
Q-score: 0.328
Zhu X, Xie X, Das H, Tan BG, Shi Y, Al-Behadili A, Peter B, Motori E, Valenzuela S, Posse V, Gustafsson CM, Hallberg BM, Falkenberg M
Cell (2022) 185 pp. 2309-2323.e24 [ DOI: doi:10.1016/j.cell.2022.05.006 Pubmed: 35662414 ]
- Polrmt (137 kDa, Cellular component from Homo sapiens)
- Dna-directed rna polymerase, mitochondrial (134 kDa, Protein from Homo sapiens)
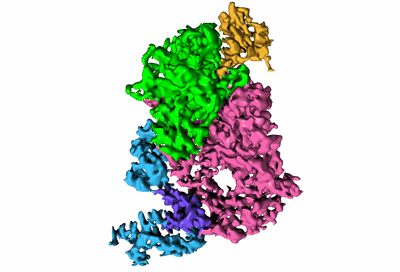
Cryo-EM structure of POLRMT in free form.
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7pzr
Q-score: 0.385
Zhu X, Xie X, Das H, Tan BG, Shi Y, Al-Behadili A, Peter B, Motori E, Valenzuela S, Posse V, Gustafsson CM, Hallberg BM, Falkenberg M
Cell (2022) 185 pp. 2309-2323.e24 [ DOI: doi:10.1016/j.cell.2022.05.006 Pubmed: 35662414 ]
- Dna-directed rna polymerase, mitochondrial (137 kDa, Protein from Homo sapiens)
- Polrmt (137 kDa, Complex from Homo sapiens)
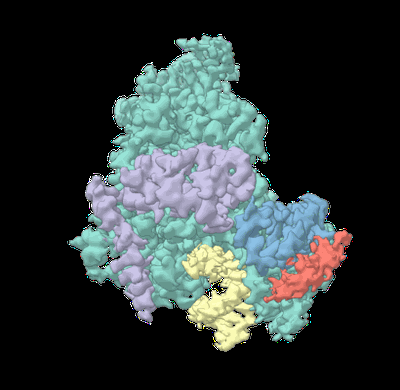
Mitochondrial DNA dependent RNA polymerase homodimer.
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7pzp
Q-score: 0.323
Zhu X, Xie X, Das H, Tan BG, Shi Y, Al-Behadili A, Peter B, Motori E, Valenzuela S, Posse V, Gustafsson CM, Hallberg BM, Falkenberg M
Cell (2022) 185 pp. 2309- [ DOI: doi:10.1016/j.cell.2022.05.006 Pubmed: 35662414 ]
- Dna-directed rna polymerase, mitochondrial (137 kDa, Protein from Homo sapiens)
- Polrmt (137 kDa, Complex)
- Mitochondrial dna-directed rna polymerase dimer. (Complex from Homo sapiens)
SARS-CoV-2 nsp12/7/8 complex with a native N-terminus nsp9
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 7thm
Q-score: 0.486
Park GJ, Osinski A, Hernandez G, Eitson JL, Majumdar A, Tonelli M, Henzler-Wildman K, Pawlowski K, Chen Z, Li Y, Schoggins JW, Tagliabracci VS
Nature (2022) 609 pp. 793-800 [ Pubmed: 35944563 DOI: doi:10.1038/s41586-022-05185-z ]
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Manganese (ii) ion (54 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Sars-cov-2 replication-transcription complex with a native n-terminus nsp9 bound to niran domain of nsp12. (172 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7aap
Q-score: 0.571
Naydenova K, Muir KW, Wu LF, Zhang Z, Coscia F, Peet MJ, Castro-Hartmann P, Qian P, Sader K, Dent K, Kimanius D, Sutherland JD, Lowe J, Barford D, Russo CJ
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2021946118 Pubmed: 33526596 ]
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*up*up*ap*ap*gp*up*up*ap*u)-3') (7 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*up*up*cp*ap*up*ap*ap*cp*up*up*ap*a)-3') (9 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 12 (110 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- [[(2~{r},3~{s},4~{r},5~{r})-5-(3-aminocarbonyl-5-fluoranyl-2-oxidanylidene-pyrazin-1-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate (529 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Nsp7-nsp8-nsp12 sars-cov2 rna-directed rna polymerase (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (Complex from Severe acute respiratory syndrome coronavirus 2)
- Nsp7-nsp8-nsp12 sars-cov2 rna-directed rna polymerase in complex with template:primer dsrna and favipiravir-rtp (Complex)
Structure of DNA Polymerase Zeta (Apo)
Resolution: 4.1 Å
EM Method: Single-particle
Fitted PDBs: 6v8p
Q-score: 0.369
Malik R, Kopylov M, Gomez-Llorente Y, Jain R, Johnson RE, Prakash L, Prakash S, Ubarretxena-Belandia I, Aggarwal AK
Nat Struct Mol Biol (2020) 27 pp. 913-924 [ DOI: doi:10.1038/s41594-020-0476-7 Pubmed: 32807989 ]
- Dna polymerase delta small subunit (55 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Dna polymerase zeta catalytic subunit (177 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Dna polymerase zeta processivity subunit (28 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Dna polymerase delta subunit 3 (40 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Apo (Complex from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Iron/sulfur cluster (351 Da, Ligand)