Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
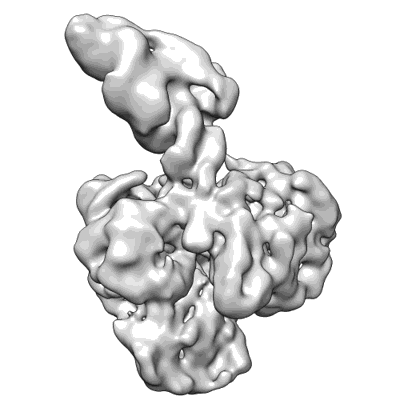
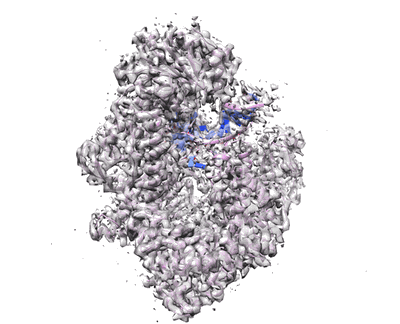
Cryo-EM structure of MLE in complex with ADP:AlF4 and U10 RNA
Resolution: 2.86 Å
EM Method: Single-particle
Fitted PDBs: 8b9g
Q-score: 0.513
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ DOI: doi:10.1016/j.molcel.2023.10.026 Pubmed: 37989319 ]
- Water (18 Da, Ligand)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Rna (5'-r(p*up*up*up*up*up*up*up*up*up*up*u)-3') (3 kDa, RNA from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
- Mle in complex with adp:alf4 and u10 rna (133 kDa, Complex from Drosophila melanogaster)
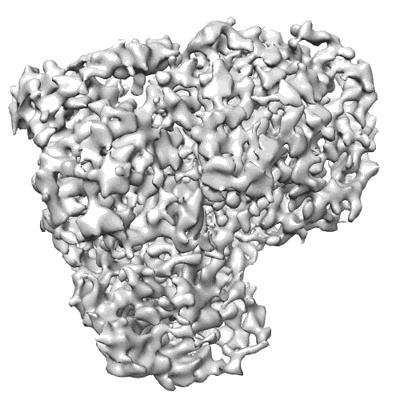
Cryo-EM structure of MLE in complex with ADP:AlF4 and UUC RNA
Resolution: 2.95 Å
EM Method: Single-particle
Fitted PDBs: 8b9i
Q-score: 0.469
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ DOI: doi:10.1016/j.molcel.2023.10.026 Pubmed: 37989319 ]
- Water (18 Da, Ligand)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Rna (5'-r(p*cp*cp*up*cp*up*up*up*cp*up*up*u)-3') (3 kDa, RNA from Drosophila melanogaster)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Mle in complex with adp:alf4 and uuc rna (133 kDa, Complex from Drosophila melanogaster)
- Magnesium ion (24 Da, Ligand)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
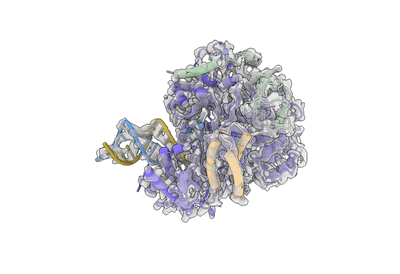
Cryo-EM structure of MLE in complex with ADP:AlF4
Resolution: 3.45 Å
EM Method: Single-particle
Fitted PDBs: 8b9j
Q-score: 0.416
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ DOI: doi:10.1016/j.molcel.2023.10.026 Pubmed: 37989319 ]
- Mle in complex with adp:alf4 (130 kDa, Cellular component from Drosophila melanogaster)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
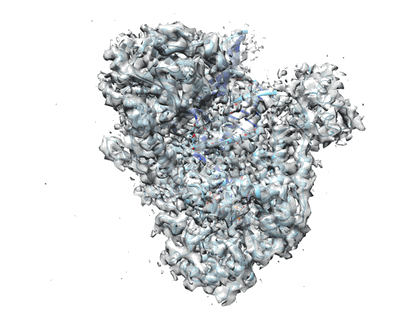
Cryo-EM structure of MLE in complex with ADP:AlF4 and SL7modUUC RNA
Resolution: 4.04 Å
EM Method: Single-particle
Fitted PDBs: 8b9k
Q-score: 0.239
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ DOI: doi:10.1016/j.molcel.2023.10.026 Pubmed: 37989319 ]
- Mle in complex with adp:alf4 and sl7moduuc rna (149 kDa, Complex from Drosophila melanogaster)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Sl7moduuc (19 kDa, RNA from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
Resolution: 3.45 Å
EM Method: Single-particle
Fitted PDBs: 8b9l
Q-score: 0.345
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ DOI: doi:10.1016/j.molcel.2023.10.026 Pubmed: 37989319 ]
- Mle (130 kDa, Cellular component from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
SARS-CoV-2 RdRp in complex with 4 Remdesivir monophosphate
Resolution: 3.89 Å
EM Method: Single-particle
Fitted PDBs: 7l1f
Q-score: 0.327
Bravo JPK, Dangerfield TL, Taylor DW, Johnson KA
Mol Cell (2021) 81 pp. 1548-1552.e4 [ DOI: doi:10.1016/j.molcel.2021.01.035 Pubmed: 33631104 ]
- Rna-directed rna polymerase (103 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*up*ap*ap*gp*ap*ap*gp*cp*up*ap*up*u*(f86)*(f86)*(f86)*(f86))-3') (5 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (7 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase, non-structural protein 8, non-structural protein 7 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (Complex)
- Rna (5'-r(p*ap*up*up*up*up*ap*ap*up*ap*gp*cp*up*up*cp*up*up*ap*g)-3') (5 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 rdrp complex with template:primer and four rmp (Complex)
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex
Resolution: 3.71 Å
EM Method: Single-particle
Fitted PDBs: 6ymw
Q-score: 0.41
De Wijngaert B, Sultana S, Singh A, Dharia C, Vanbuel H, Shen J, Vasilchuk D, Martinez SE, Kandiah E, Patel SS, Das K
Mol Cell (2021) 81 pp. 268- [ DOI: doi:10.1016/j.molcel.2020.11.016 Pubmed: 33278362 ]
- Dna (Complex from Saccharomyces cerevisiae)
- Mitochondrial transcription factor 1 (41 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondria dna-dependent rna polymerase pre-initiation complex (202 kDa, Complex)
- Chains: t (10 kDa, DNA from Saccharomyces cerevisiae S288C)
- Mitochondria dna-dependent rna polymerase pre-initiation complex (Complex from Saccharomyces cerevisiae)
- Dna-directed rna polymerase, mitochondrial (143 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- [[(2~{r},3~{s},4~{r},5~{r})-5-[2,4-bis(oxidanylidene)pyrimidin-1-yl]-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-sulfanyl-phosphoryl] phosphono hydrogen phosphate (500 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rna (pppgpg) (805 Da, RNA from synthetic construct)
- Chains: n (10 kDa, DNA from Saccharomyces cerevisiae S288C)
- Rna (pppgpg) (Complex from synthetic construct)
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription pre-initiation complex
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 6ymv
Q-score: 0.536
De Wijngaert B, Sultana S, Singh A, Dharia C, Vanbuel H, Shen J, Vasilchuk D, Martinez SE, Kandiah E, Patel SS, Das K
Mol Cell (2021) 81 pp. 268- [ DOI: doi:10.1016/j.molcel.2020.11.016 Pubmed: 33278362 ]
- Dna (Complex from Saccharomyces cerevisiae)
- Mitochondrial transcription factor 1 (41 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondria dna-dependent rna polymerase pre-initiation complex (202 kDa, Complex)
- Dna (33-mer) non-template (10 kDa, DNA from Saccharomyces cerevisiae)
- Dna-directed rna polymerase, mitochondrial (143 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Dna (33-mer) template (10 kDa, DNA from Saccharomyces cerevisiae)
- Mitochondria dna-dependent rna polymerase pre-initiation complex (Complex from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
Electron cryo-microscopy of a transcribing Pol IIp-Capping Enzyme complex
Martinez-Rucobo FW, Kohler R, van de Waterbeemd M, Heck AJR, Hemann M, Herzog F, Stark H, Cramer P
Mol Cell (2015) 58 pp. 1079-1089 [ DOI: doi:10.1016/j.molcel.2015.04.004 Pubmed: 25959396 ]
- Mrna-capping enzyme (180 kDa, Protein from Saccharomyces cerevisiae)
- Transcribing pol iip-capping enzyme complex from yeast (720 kDa, Sample)
- Dna-directed rna polymerase ii (520 kDa, Protein from Saccharomyces cerevisiae)
- Dna-rna scaffold (20 kDa, Macromolecule from Saccharomyces cerevisiae)
Electron cryo-microscopy of a transcribing Pol IIp complex
Martinez-Rucobo FW, Kohler R, van de Waterbeemd M, Heck AJR, Hemann M, Herzog F, Stark H, Cramer P
Mol Cell (2015) 58 pp. 1079-1089 [ DOI: doi:10.1016/j.molcel.2015.04.004 Pubmed: 25959396 ]
- Dna-rna scaffold (20 kDa, Macromolecule from Saccharomyces cerevisiae)
- Dna-directed rna polymerase ii (520 kDa, Protein from Saccharomyces cerevisiae)
- Transcribing pol iip complex from yeast (540 kDa, Sample)