Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
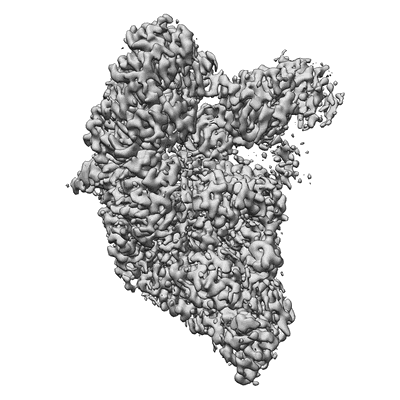
Structure of SARS-CoV-2 backtracked complex bound to nsp13 helicase - nsp13(1)-BTC
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7krn
Q-score: 0.476
Malone B, Chen J, Wang Q, Llewellyn E, Choi YJ, Olinares PDB, Cao X, Hernandez C, Eng ET, Chait BT, Shaw DE, Landick R, Darst SA, Campbell EA
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2102516118 Pubmed: 33883267 ]
- Rna (43-mer) (17 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (37-mer) (12 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Sars-cov-2 backtracked complex bound to nsp13 helicase - nsp13(1)-btc (Complex from Severe acute respiratory syndrome coronavirus 2)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
Structure of SARS-CoV-2 backtracked complex complex bound to nsp13 helicase - nsp13(2)-BTC
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7kro
Q-score: 0.416
Malone B, Chen J, Wang Q, Llewellyn E, Choi YJ, Olinares PDB, Cao X, Hernandez C, Eng ET, Chait BT, Shaw DE, Landick R, Darst SA, Campbell EA
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2102516118 Pubmed: 33883267 ]
- Rna (43-mer) (17 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (37-mer) (12 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 backtracked complex complex bound to nsp13 helicase - nsp13(2)-btc (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
Structure of SARS-CoV-2 backtracked complex complex bound to nsp13 helicase - BTC (local refinement)
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 7krp
Q-score: 0.361
Malone B, Chen J, Wang Q, Llewellyn E, Choi YJ, Olinares PDB, Cao X, Hernandez C, Eng ET, Chait BT, Shaw DE, Landick R, Darst SA, Campbell EA
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2102516118 Pubmed: 33883267 ]
- Rna (37-mer) (12 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 backtracked complex complex bound to nsp13 helicase - btc (local refinement) (Complex from Severe acute respiratory syndrome coronavirus 2)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (36-mer) (17 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)