Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
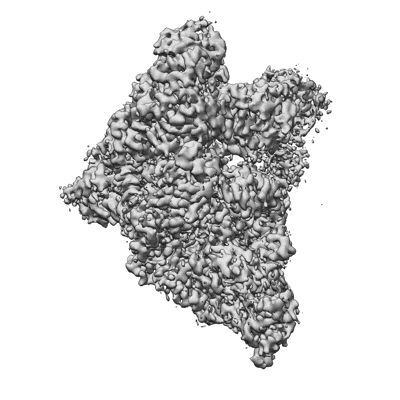
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 7uo4
Q-score: 0.457
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ DOI: doi:10.1038/s41586-022-05664-3 Pubmed: 36725929 ]
- Water (18 Da, Ligand)
- [[(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate (531 Da, Ligand)
- Sars-cov-2 replication-transcription complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
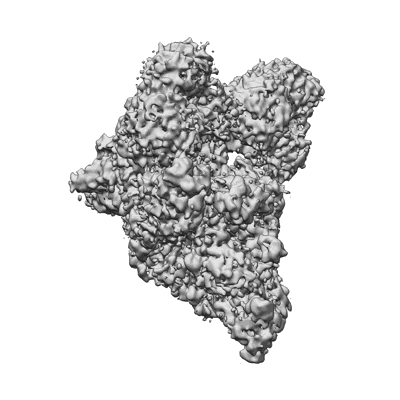
SARS-CoV-2 replication-transcription complex bound to ATP, in a pre-catalytic state
Resolution: 3.09 Å
EM Method: Single-particle
Fitted PDBs: 7uo7
Q-score: 0.451
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ DOI: doi:10.1038/s41586-022-05664-3 Pubmed: 36725929 ]
- Sars-cov-2 replication-transcription complex + atp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
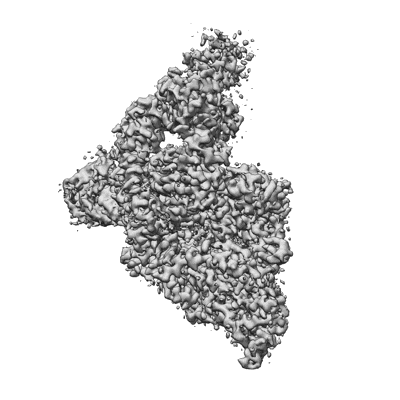
SARS-CoV-2 replication-transcription complex bound to UTP, in a pre-catalytic state
Resolution: 3.13 Å
EM Method: Single-particle
Fitted PDBs: 7uo9
Q-score: 0.487
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ DOI: doi:10.1038/s41586-022-05664-3 Pubmed: 36725929 ]
- Water (18 Da, Ligand)
- Sars-cov-2 replication-transcription complex + utp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- Uridine 5'-triphosphate (484 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
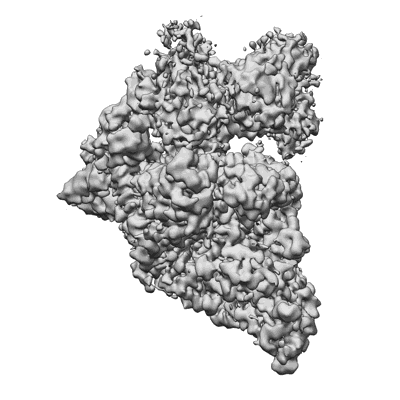
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - engaged class
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7rdy
Q-score: 0.472
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Sars-cov-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-rtc - engaged class (Complex from Severe acute respiratory syndrome coronavirus 2)
- Template rna (17 kDa, RNA from synthetic construct)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - swiveled class
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7re0
Q-score: 0.323
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Template rna (17 kDa, RNA from synthetic construct)
- Sars-cov-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-rtc - swiveled class (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(1)-RTC
Resolution: 3.17 Å
EM Method: Single-particle
Fitted PDBs: 7re2
Q-score: 0.464
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Sars-cov-2 replication/transcription complex bound to nsp13 helicase - nsp13(1)-rtc (Complex from Severe acute respiratory syndrome coronavirus 2)
- Template rna (17 kDa, RNA from synthetic construct)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - open class
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7rdx
Q-score: 0.442
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Template rna (17 kDa, RNA from synthetic construct)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 replication/transcription complex bound to nsp13 helicase - nsp13(2)-rtc - open class (Complex from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC - apo class
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7rdz
Q-score: 0.31
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Template rna (17 kDa, RNA from synthetic construct)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 replication/transcription complex bound to nsp13 helicase - nsp13(2)-rtc - apo class (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
SARS-CoV-2 replication-transcription complex bound to nsp13 helicase - nsp13(2)-RTC dimer
Resolution: 3.33 Å
EM Method: Single-particle
Fitted PDBs: 7re3
Q-score: 0.321
Chen J, Wang Q, Malone B, Llewellyn E, Pechersky Y, Maruthi K, Eng ET, Perry JK, Campbell EA, Shaw DE, Darst SA
Nat Struct Mol Biol (2022) 29 pp. 250-260 [ DOI: doi:10.1038/s41594-022-00734-6 Pubmed: 35260847 ]
- Template rna (17 kDa, RNA from synthetic construct)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Product rna (11 kDa, RNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Sars-cov-2 replication/transcription complex bound to nsp13 helicase - nsp13(2)-rtc dimer (Complex from Severe acute respiratory syndrome coronavirus 2)
Structure of SARS-CoV-2 backtracked complex bound to nsp13 helicase - nsp13(1)-BTC
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7krn
Q-score: 0.476
Malone B, Chen J, Wang Q, Llewellyn E, Choi YJ, Olinares PDB, Cao X, Hernandez C, Eng ET, Chait BT, Shaw DE, Landick R, Darst SA, Campbell EA
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2102516118 Pubmed: 33883267 ]
- Rna (43-mer) (17 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (37-mer) (12 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Chapso (631 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (67 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Aluminum fluoride (83 Da, Ligand)
- Sars-cov-2 backtracked complex bound to nsp13 helicase - nsp13(1)-btc (Complex from Severe acute respiratory syndrome coronavirus 2)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)