Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
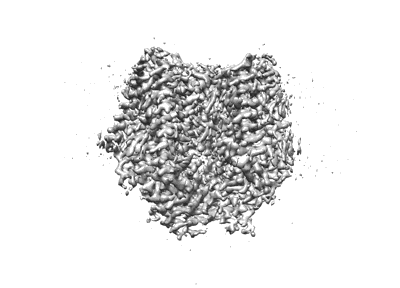
cryo-EM structure of apo Clostridioides difficile toxin B
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8qen
Q-score: 0.452
Kinsolving J, Bous J, Kozielewicz P, Kosenina S, Shekhani R, Gratz L, Masuyer G, Wang Y, Stenmark P, Dong M, Schulte G
Cell Rep (2024) 43 pp. 113727-113727 [ Pubmed: 38308843 DOI: doi:10.1016/j.celrep.2024.113727 ]
- Apo-structure of clostridioides difficile toxin b (Cellular component from Priestia megaterium DSM 319)
- Toxin b (273 kDa, Protein from Clostridioides difficile)
- Masked region 2 (Cellular component from Priestia megaterium DSM 319)
- Masked region 1 (Cellular component from Priestia megaterium DSM 319)
- Zinc ion (65 Da, Ligand)
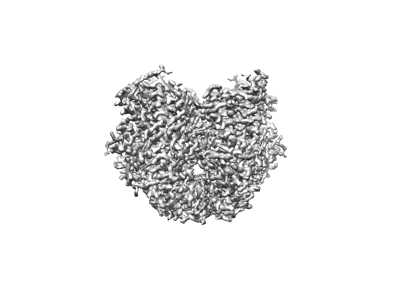
cryo-EM structure complex of Frizzled-7 and Clostridioides difficile toxin B
Resolution: 3.26 Å
EM Method: Single-particle
Fitted PDBs: 8qeo
Q-score: 0.416
Kinsolving J, Bous J, Kozielewicz P, Kosenina S, Shekhani R, Gratz L, Masuyer G, Wang Y, Stenmark P, Dong M, Schulte G
Cell Rep (2024) 43 pp. 113727-113727 [ Pubmed: 38308843 DOI: doi:10.1016/j.celrep.2024.113727 ]
- Masked region--crop domain (Cellular component from Priestia megaterium DSM 319)
- Toxin b (273 kDa, Protein from Clostridioides difficile)
- Masked region-core domain (Cellular component from Priestia megaterium DSM 319)
- Masked region-fzd7 crd and leg of tcdb (Cellular component from Priestia megaterium DSM 319)
- Complex of frizzled-7 and clostridioides difficile toxin b (Complex from Spodoptera frugiperda)
- Zinc ion (65 Da, Ligand)
- Frizzled-7 (67 kDa, Protein from Homo sapiens)
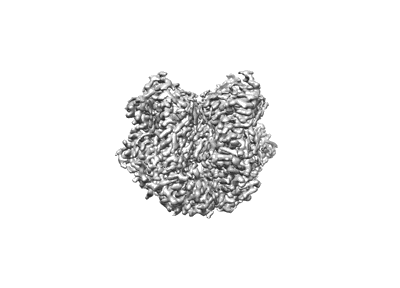
Structure of the wild-type Arabidopsis ABCB19 in the apo state
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8woi
Q-score: 0.49
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ DOI: doi:10.1126/science.adj4591 Pubmed: 38513023 ]
Structure of the wild-type Arabidopsis ABCB19 in the brassinolide-bound state
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8wom
Q-score: 0.451
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ DOI: doi:10.1126/science.adj4591 Pubmed: 38513023 ]
Structure of the wild-type Arabidopsis ABCB19 in the brassinolide and AMP-PNP bound state
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 8woo
Q-score: 0.401
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ DOI: doi:10.1126/science.adj4591 Pubmed: 38513023 ]
- Purified wild-type abcb19 protein in complex with brassinolide and amp-pnp (Cellular component from Arabidopsis thaliana)
- Magnesium ion (24 Da, Ligand)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Abc transporter b family member 19 (136 kDa, Protein from Arabidopsis thaliana)
- Brassinolide (480 Da, Ligand)
Structure of the Arabidopsis E529Q/E1174Q ABCB19 in the ATP bound state
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8wp0
Q-score: 0.464
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ DOI: doi:10.1126/science.adj4591 Pubmed: 38513023 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Purified e529q/e1174q mutant abcb19 protein in complex with atp (Cellular component from Arabidopsis thaliana)
- Abc transporter b family member 19 (136 kDa, Protein from Arabidopsis thaliana)
The cryo-EM structure of insect gustatory receptor Gr43a from Drosophila melanogaster
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8jm9
Q-score: 0.593
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8jma
Q-score: 0.637
The cryo-EM structure of insect gustatory receptor Gr64a from Drosophila melanogaster
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8jme
Q-score: 0.637
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8jmh
Q-score: 0.615