Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Cryo-EM structure of the ClpP degradation system in Streptomyces hawaiiensis
Resolution: 2.34 Å
EM Method: Single-particle
Fitted PDBs: 8xn4
Q-score: 0.56
Cryo-EM structure of the ClpC1:ClpP1P2 degradation complex in Streptomyces hawaiiensis
Resolution: 1.96 Å
EM Method: Single-particle
Fitted PDBs: 8xon
Q-score: 0.503
Xu X, Wang Y, Huang W, Li D, Deng Z, Long F
Mbio (2024) 15 pp. e0003124-e0003124 [ DOI: doi:10.1128/mbio.00031-24 Pubmed: 38501868 ]
- Atp-dependent clp protease proteolytic subunit (22 kDa, Protein from Streptomyces hawaiiensis)
- Casein (2 kDa, Protein from Bos taurus)
- Atp-dependent clp protease proteolytic subunit (24 kDa, Protein from Streptomyces hawaiiensis)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Clpp1,clpp2,clpc1,casein (Complex from Streptomyces hawaiiensis)
- Ndp-hexose 4-ketoreductase (77 kDa, Protein from Streptomyces hawaiiensis)
Cryo-EM structure of the ClpC1:ClpP1P2 degradation complex in Streptomyces hawaiiensis
Resolution: 1.84 Å
EM Method: Single-particle
Fitted PDBs: 8xoo
Q-score: 0.514
Xu X, Wang Y, Huang W, Li D, Deng Z, Long F
Mbio (2024) 15 pp. e0003124-e0003124 [ DOI: doi:10.1128/mbio.00031-24 Pubmed: 38501868 ]
- Clpp1,clpp2,clpc1,casein (Complex from Streptomyces hawaiiensis)
- Atp-dependent clp protease proteolytic subunit (22 kDa, Protein from Streptomyces hawaiiensis)
- Atp-dependent clp protease proteolytic subunit (24 kDa, Protein from Streptomyces hawaiiensis)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Casein (2 kDa, Protein from Bos taurus)
- Ndp-hexose 4-ketoreductase (77 kDa, Protein from Streptomyces hawaiiensis)
Cryo-EM structure of ClpP1P2 in complex with ADEP1 from Streptomyces hawaiiensis
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8xop
Q-score: 0.498
Xu X, Wang Y, Huang W, Li D, Deng Z, Long F
Mbio (2024) 15 pp. e0003124-e0003124 [ DOI: doi:10.1128/mbio.00031-24 Pubmed: 38501868 ]
- Adep1 (736 Da, Protein from synthetic construct)
- Atp-dependent clp protease proteolytic subunit (22 kDa, Protein from Streptomyces hawaiiensis)
- Atp-dependent clp protease proteolytic subunit (24 kDa, Protein from Streptomyces hawaiiensis)
- Clpp1,clpp2 (Complex from Streptomyces hawaiiensis)
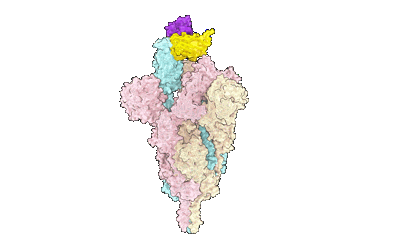
SARS-CoV-2 Omicron variant spike in complex with Fab 9A8 (State 1)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7wt7
Q-score: 0.385
Liu S, Jia Z, Nie J, Liang Z, Xie J, Wang L, Zhang L, Wang X, Wang Y, Huang W
Cell Discov (2022) 8 pp. 81-81 [ Pubmed: 35977939 DOI: doi:10.1038/s41421-022-00443-w ]
- Sars-cov-2 omicron variant spike in complex with fab 9a8 (state 1) (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Heavy chain of fab 9a8 (12 kDa, Protein from Homo sapiens)
- Light chain of fab 9a8 (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
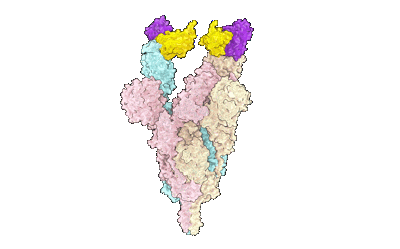
SARS-CoV-2 Omicron variant spike in complex with Fab 9A8 (State 2)
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7wt8
Q-score: 0.358
Liu S, Jia Z, Nie J, Liang Z, Xie J, Wang L, Zhang L, Wang X, Wang Y, Huang W
Cell Discov (2022) 8 pp. 81-81 [ Pubmed: 35977939 DOI: doi:10.1038/s41421-022-00443-w ]
- Sars-cov-2 omicron variant spike in complex with fab 9a8 (state 2) (Complex from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron variant spike (Complex from Homo sapiens)
- Fab 9a8 (Complex)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Heavy chain of fab 9a8 (12 kDa, Protein from Homo sapiens)
- Light chain of fab 9a8 (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
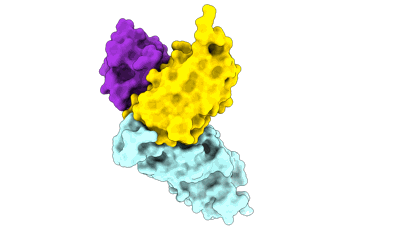
SARS-CoV-2 Omicron variant spike RBD in complex with Fab 9A8
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 7wt9
Q-score: 0.336
Liu S, Jia Z, Nie J, Liang Z, Xie J, Wang L, Zhang L, Wang X, Wang Y, Huang W
Cell Discov (2022) 8 pp. 81-81 [ Pubmed: 35977939 DOI: doi:10.1038/s41421-022-00443-w ]
- Heavy chain of fab 9a8 (12 kDa, Protein from Homo sapiens)
- Sars-cov-2 omicron variant spike (Complex)
- Light chain of fab 9a8 (11 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron variant spike rbd in complex with fab 9a8 (Complex from Homo sapiens)
- Fab 9a8 (Complex from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 spike glycoprotein trimer in closed state
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 7wgv
Q-score: 0.513
Zhao MM, Zhu Y, Zhang L, Zhong G, Tai L, Liu S, Yin G, Lu J, He Q, Li MJ, Zhao RX, Wang H, Huang W, Fan C, Shuai L, Wen Z, Wang C, He X, Chen Q, Liu B, Xiong X, Bu Z, Wang Y, Sun F, Yang JK
Cell Discov (2022) 8 pp. 53- [ DOI: doi:10.1038/s41421-022-00419-w ]
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 b.1.1.7 spike glycoprotein trimer (Complex from Severe acute respiratory syndrome coronavirus 2)
- Linoleic acid (280 Da, Ligand)
- 3-[5-[(z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1h-pyrrol-2-yl]methyl]-4-methyl-1h-pyrrol-3-yl]propanoic acid (584 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 spike glycoprotein trimer in closed state after treatment with Cathepsin L
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7wgx
Q-score: 0.468
Zhao MM, Zhu Y, Zhang L, Zhong G, Tai L, Liu S, Yin G, Lu J, He Q, Li MJ, Zhao RX, Wang H, Huang W, Fan C, Shuai L, Wen Z, Wang C, He X, Chen Q, Liu B, Xiong X, Bu Z, Wang Y, Sun F, Yang JK
Cell Discov (2022) 8 pp. 53- [ DOI: doi:10.1038/s41421-022-00419-w ]
- 3-[5-[(z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1h-pyrrol-2-yl]methyl]-4-methyl-1h-pyrrol-3-yl]propanoic acid (584 Da, Ligand)
- Sars-cov-2 b.1.1.7 spike glycoprotein trimer (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 spike glycoprotein trimer in Intermediate state
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 7wgy
Q-score: 0.354