Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
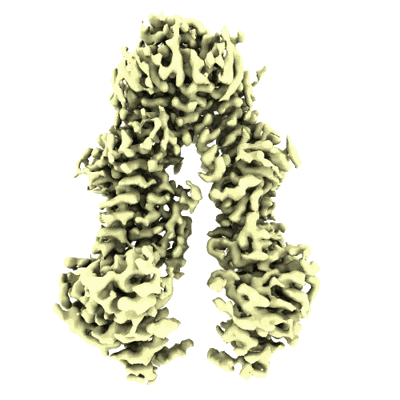
Structure of the wild-type Arabidopsis ABCB19 in the apo state
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8woi
Q-score: 0.49
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ Pubmed: 38513023 DOI: doi:10.1126/science.adj4591 ]
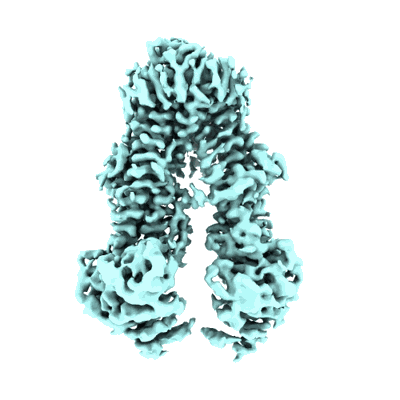
Structure of the wild-type Arabidopsis ABCB19 in the brassinolide-bound state
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8wom
Q-score: 0.451
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ Pubmed: 38513023 DOI: doi:10.1126/science.adj4591 ]
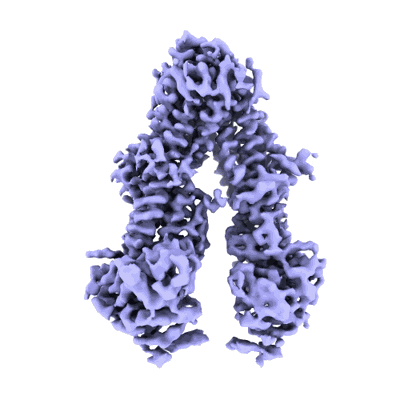
Structure of the wild-type Arabidopsis ABCB19 in the brassinolide and AMP-PNP bound state
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 8woo
Q-score: 0.401
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ Pubmed: 38513023 DOI: doi:10.1126/science.adj4591 ]
- Brassinolide (480 Da, Ligand)
- Purified wild-type abcb19 protein in complex with brassinolide and amp-pnp (Cellular component from Arabidopsis thaliana)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Abc transporter b family member 19 (136 kDa, Protein from Arabidopsis thaliana)
- Magnesium ion (24 Da, Ligand)
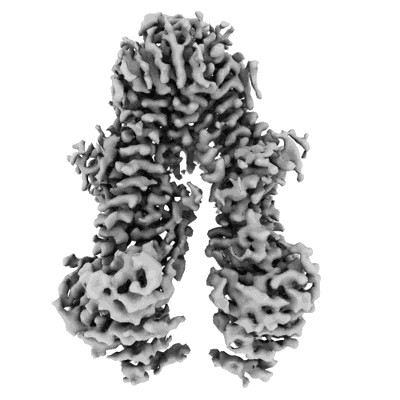
Structure of the Arabidopsis E529Q/E1174Q ABCB19 in the ATP bound state
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8wp0
Q-score: 0.464
Ying W, Wang Y, Wei H, Luo Y, Ma Q, Zhu H, Janssens H, Vukasinovic N, Kvasnica M, Winne JM, Gao Y, Tan S, Friml J, Liu X, Russinova E, Sun L
Science (2024) 383 pp. eadj4591-eadj4591 [ Pubmed: 38513023 DOI: doi:10.1126/science.adj4591 ]
- Magnesium ion (24 Da, Ligand)
- Purified e529q/e1174q mutant abcb19 protein in complex with atp (Cellular component from Arabidopsis thaliana)
- Abc transporter b family member 19 (136 kDa, Protein from Arabidopsis thaliana)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Resolution: 3.66 Å
EM Method: Single-particle
Fitted PDBs: 8k45
Q-score: 0.408
Yao H, Wang H, Zhang Z, Lu Y, Zhang Z, Zhang Y, Xiong X, Wang Y, Wang Z, Yang H, Zhao J, Xu W
MedComm (2020) (2023) 4 pp. e397-e397 [ Pubmed: 37901798 DOI: doi:10.1002/mco2.397 ]
- Nb4 nanobody (13 kDa, Protein from Vicugna pacos)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 omicron ba.1 spike glycoprotein complex with nb4 nanobody (Complex from Severe acute respiratory syndrome coronavirus 2)
Resolution: 3.37 Å
EM Method: Single-particle
Fitted PDBs: 8k46
Q-score: 0.377
Yao H, Wang H, Zhang Z, Lu Y, Zhang Z, Zhang Y, Xiong X, Wang Y, Wang Z, Yang H, Zhao J, Xu W
MedComm (2020) (2023) 4 pp. e397-e397 [ Pubmed: 37901798 DOI: doi:10.1002/mco2.397 ]
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Nanobody nb4 (13 kDa, Protein from Vicugna pacos)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 omicron ba.1 spike glycoprotein complex with nb4 nanobody (Complex from Severe acute respiratory syndrome coronavirus 2)
Resolution: 3.54 Å
EM Method: Single-particle
Fitted PDBs: 8k47
Q-score: 0.334
Yao H, Wang H, Zhang Z, Lu Y, Zhang Z, Zhang Y, Xiong X, Wang Y, Wang Z, Yang H, Zhao J, Xu W
MedComm (2020) (2023) 4 pp. e397-e397 [ Pubmed: 37901798 DOI: doi:10.1002/mco2.397 ]
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Nanobody nb4 (13 kDa, Protein from Vicugna pacos)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 omicron ba.1 spike glycoprotein complex with nb4 nanobody (Complex from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM Structure of SARS-CoV-2 BA.2 Spike protein in complex with BA7535
Resolution: 2.44 Å
EM Method: Single-particle
Fitted PDBs: 8h7l
Q-score: 0.377
Wang Y, Yan A, Song D, Duan M, Dong C, Chen J, Jiang Z, Gao Y, Rao M, Feng J, Zhang Z, Qi R, Ma X, Liu H, Yu B, Wang Q, Zong M, Jiao J, Xing P, Pan R, Li D, Xiao J, Sun J, Li Y, Zhang L, Shen Z, Sun B, Zhao Y, Zhang L, Dai J, Zhao J, Wang L, Dou C, Liu Z, Zhao J
Nat Commun (2024) 15 pp. 842-842 [ Pubmed: 38287016 DOI: doi:10.1038/s41467-024-45050-3 ]
- Ba7535 fab light chain (23 kDa, Protein from Homo sapiens)
- Ba7535 fab heavt chain (49 kDa, Protein from Homo sapiens)
- Spike glycoprotein (124 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 ba.2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Cryo-em structure of sars-cov-2 ba.2 spike protein in complex with ba7535 (Complex)
- Ba7535 (Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Cryo-EM structure of SARS-CoV-2 BA.2 RBD in complex with BA7535 fab (local refinement)
Resolution: 3.07 Å
EM Method: Single-particle
Fitted PDBs: 8h7z
Q-score: 0.461
Wang Y, Yan A, Song D, Duan M, Dong C, Chen J, Jiang Z, Gao Y, Rao M, Feng J, Zhang Z, Qi R, Ma X, Liu H, Yu B, Wang Q, Zong M, Jiao J, Xing P, Pan R, Li D, Xiao J, Sun J, Li Y, Zhang L, Shen Z, Sun B, Zhao Y, Zhang L, Dai J, Zhao J, Wang L, Dou C, Liu Z, Zhao J
Nat Commun (2024) 15 pp. 842-842 [ Pubmed: 38287016 DOI: doi:10.1038/s41467-024-45050-3 ]
- Ba7535 fab (23 kDa, Protein from Homo sapiens)
- Ba7535 fab (49 kDa, Protein from Homo sapiens)
- Spike glycoprotein (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Cryo-em structure of sars-cov-2 ba.2 rbd in complex with ba7535 fab (local refinement) (Complex from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of Arabidopsis thaliana SOS1 in an occluded state
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7y3e
Q-score: 0.539
Wang Y, Pan C, Chen Q, Xie Q, Gao Y, He L, Li Y, Dong Y, Jiang X, Zhao Y
Nat Commun (2023) 14 pp. 4487-4487 [ Pubmed: 37495621 DOI: doi:10.1038/s41467-023-40215-y ]
- Sodium/hydrogen exchanger 7 (127 kDa, Protein from Arabidopsis thaliana)
- Arabidopsis sodium/hydrogen exchanger 7 (sos1), homodimer (Complex from Arabidopsis thaliana)
- Hexadecane (226 Da, Ligand)