Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
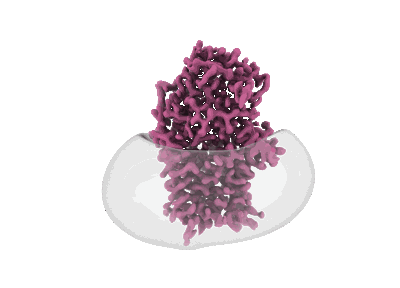
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8b0k
Q-score: 0.541
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
Resolution: 3.13 Å
EM Method: Single-particle
Fitted PDBs: 8b0l
Q-score: 0.511
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 8b0m
Q-score: 0.552
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
Resolution: 2.67 Å
EM Method: Single-particle
Fitted PDBs: 8b0n
Q-score: 0.6
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
Resolution: 3.02 Å
EM Method: Single-particle
Fitted PDBs: 8b0o
Q-score: 0.534
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
- Apolipoprotein n-acyltransferase (59 kDa, Protein from Escherichia coli K-12)
- Apolipoprotein n-acyltransferase (Cellular component from Escherichia coli K-12)
- [(2~{r})-3-[(2~{r})-3-[[(2~{r})-1-[[(2~{r})-1-[[(2~{r})-6-[(2-aminophenyl)carbonylamino]-1-azanyl-1-oxidanylidene-hexan-2-yl]amino]-3-oxidanyl-1-oxidanylidene-propan-2-yl]amino]-3-oxidanyl-1-oxidanylidene-propan-2-yl]amino]-2-(hexadecanoylamino)-3-oxidanylidene-propyl]sulfanyl-2-hexadecanoyloxy-propyl] hexadecanoate (1 kDa, Ligand)
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3
Resolution: 2.86 Å
EM Method: Single-particle
Fitted PDBs: 8b0p
Q-score: 0.564
Smithers L, Degtjarik O, Weichert D, Huang CY, Boland C, Bowen K, Oluwole A, Lutomski C, Robinson CV, Scanlan EM, Wang M, Olieric V, Shalev-Benami M, Caffrey M
Sci Adv (2023) 9 pp. eadf5799-eadf5799 [ Pubmed: 37390210 DOI: doi:10.1126/sciadv.adf5799 ]
- [(2~{s})-3-[(2~{s})-3-azanyl-2-(hexadecanoylamino)-3-oxidanylidene-propyl]sulfanyl-2-hexadecanoyloxy-propyl] hexadecanoate (909 Da, Ligand)
- Apolipoprotein n-acyltransferase (Cellular component from Escherichia coli K-12)
- Apolipoprotein n-acyltransferase (59 kDa, Protein from Escherichia coli K-12)
- Pam3-skkkk (621 Da, Protein from synthetic construct)