Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
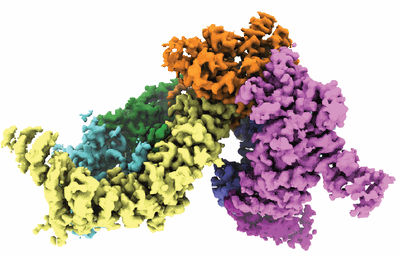
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Resolution: 4.9 Å
EM Method: Single-particle
Fitted PDBs: 7o3w
Q-score: 0.169
Gupta TK, Klumpe S, Gries K, Heinz S, Wietrzynski W, Ohnishi N, Niemeyer J, Spaniol B, Schaffer M, Rast A, Ostermeier M, Strauss M, Plitzko JM, Baumeister W, Rudack T, Sakamoto W, Nickelsen J, Schuller JM, Schroda M, Engel BD
Cell (2021) 184 pp. 3643- [ Pubmed: 34166613 DOI: doi:10.1016/j.cell.2021.05.011 ]
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7o3x
Q-score: 0.213
Gupta TK, Klumpe S, Gries K, Heinz S, Wietrzynski W, Ohnishi N, Niemeyer J, Spaniol B, Schaffer M, Rast A, Ostermeier M, Strauss M, Plitzko JM, Baumeister W, Rudack T, Sakamoto W, Nickelsen J, Schuller JM, Schroda M, Engel BD
Cell (2021) 184 pp. 3643- [ Pubmed: 34166613 DOI: doi:10.1016/j.cell.2021.05.011 ]
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 7o3y
Q-score: 0.208
Gupta TK, Klumpe S, Gries K, Heinz S, Wietrzynski W, Ohnishi N, Niemeyer J, Spaniol B, Schaffer M, Rast A, Ostermeier M, Strauss M, Plitzko JM, Baumeister W, Rudack T, Sakamoto W, Nickelsen J, Schuller JM, Schroda M, Engel BD
Cell (2021) 184 pp. 3643- [ Pubmed: 34166613 DOI: doi:10.1016/j.cell.2021.05.011 ]
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 7o40
Q-score: 0.186
Gupta TK, Klumpe S, Gries K, Heinz S, Wietrzynski W, Ohnishi N, Niemeyer J, Spaniol B, Schaffer M, Rast A, Ostermeier M, Strauss M, Plitzko JM, Baumeister W, Rudack T, Sakamoto W, Nickelsen J, Schuller JM, Schroda M, Engel BD
Cell (2021) 184 pp. 3643- [ Pubmed: 34166613 DOI: doi:10.1016/j.cell.2021.05.011 ]
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Resolution: 5.0 Å
EM Method: Single-particle
Fitted PDBs: 7o3z
Q-score: 0.168
Gupta TK, Klumpe S, Gries K, Heinz S, Wietrzynski W, Ohnishi N, Niemeyer J, Spaniol B, Schaffer M, Rast A, Ostermeier M, Strauss M, Plitzko JM, Baumeister W, Rudack T, Sakamoto W, Nickelsen J, Schuller JM, Schroda M, Engel BD
Cell (2021) 184 pp. 3643- [ Pubmed: 34166613 DOI: doi:10.1016/j.cell.2021.05.011 ]
Structure of the SMG1-SMG8-SMG9 complex
Resolution: 3.45 Å
EM Method: Single-particle
Fitted PDBs: 6syt
Q-score: 0.492
Gat Y, Schuller JM, Lingaraju M, Weyher E, Bonneau F, Strauss M, Murray PJ, Conti E
Nat Struct Mol Biol (2019) 26 pp. 1089-1093 [ Pubmed: 31792449 DOI: doi:10.1038/s41594-019-0342-7 ]
- Inositol hexakisphosphate (660 Da, Ligand)
- Smg1-smg8-smg9 pikk kinase complex (Complex from Homo sapiens)
- Protein smg9 (57 kDa, Protein from Homo sapiens)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Protein smg8 (109 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Smg1,serine/threonine-protein kinase smg1,smg1,serine/threonine-protein kinase smg1,smg1,serine/threonine-protein kinase smg1,smg1,serine/threonine-protein kinase smg1,smg1,serine/threonine-protein kinase smg1 (401 kDa, Protein from Homo sapiens)
Cryo-EM structure of a poly(A) RNP bound to the Pan2-Pan3 deadenylase
Resolution: 4.8 Å
EM Method: Single-particle
Fitted PDBs: 6r5k
Q-score: 0.197
Schafer IB, Yamashita M, Schuller JM, Schussler S, Reichelt P, Strauss M, Conti E
Cell (2019) 177 pp. 1619- [ DOI: doi:10.1016/j.cell.2019.04.013 Pubmed: 31104843 ]
- Pan2-pan3 deadenylation complex subunit pan3 (53 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Pan2-pan3 deadenylation complex catalytic subunit pan2 (127 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Polyadenylate-binding protein, cytoplasmic and nuclear (64 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Ternary complex of pan2-pan3, pab1 and poly(a) rna (498 kDa, Complex)
- Rna (Complex from Saccharomyces cerevisiae S288C)
- Pan2-pan3 deadenylation complex subunit pan3 (77 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Polyadenylate-binding protein, cytoplasmic and nuclear (Complex from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Magnesium ion (24 Da, Ligand)
- Pan2-pan3 (Complex from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Poly(a) rna (29 kDa, RNA from Saccharomyces cerevisiae S288C)