Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
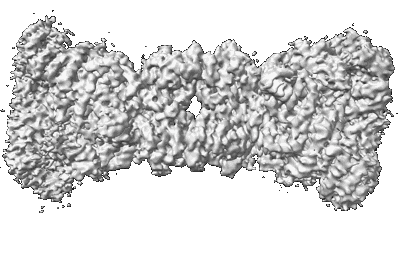
Yeast U1 snRNP with humanized U1C Zinc-Finger domain
Resolution: 3.49 Å
EM Method: Single-particle
Fitted PDBs: 8w2o
Q-score: 0.417
Chalivendra S, Shi S, Li X, Kuang Z, Giovinazzo J, Zhang L, Rossi J, Wang J, Saviola A, Welty R, Liu S, Vaeth K, Zhou ZH, Hansen K, Taliaferro JM, Zhao R
RNA (2024) [ DOI: doi:10.1261/rna.079917.123 Pubmed: 38688558 ]
- Protein luc7 (30 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1 small nuclear ribonucleoprotein 70 kda homolog (34 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1snrnp (Complex from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein sm d1 (16 kDa, Protein from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein sm d3 (11 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1 small nuclear ribonucleoprotein component snu71 (5 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1 small nuclear ribonucleoprotein component prp42 (65 kDa, Protein from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein-associated protein b (22 kDa, Protein from Saccharomyces cerevisiae S288C)
- Act1 pre-mrna (59 kDa, RNA from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein g (8 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1 small nuclear ribonucleoprotein a (40 kDa, Protein from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein sm d2 (12 kDa, Protein from Saccharomyces cerevisiae S288C)
- U1 snrna (182 kDa, RNA from Saccharomyces cerevisiae S288C)
- U1 small nuclear ribonucleoprotein c (27 kDa, Protein from Saccharomyces cerevisiae S288C)
- Pre-mrna-processing factor 39 (74 kDa, Protein from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein f (9 kDa, Protein from Saccharomyces cerevisiae S288C)
- Protein nam8 (57 kDa, Protein from Saccharomyces cerevisiae S288C)
- 56 kda u1 small nuclear ribonucleoprotein component (56 kDa, Protein from Saccharomyces cerevisiae S288C)
- Small nuclear ribonucleoprotein e (10 kDa, Protein from Saccharomyces cerevisiae S288C)
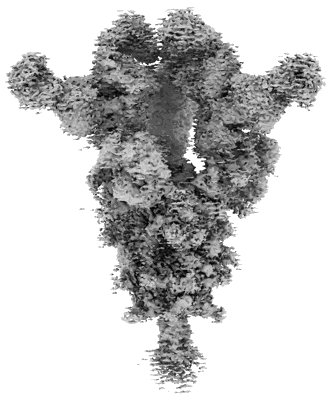
Cryo-EM structure of SARS-CoV-2 Omicron BA.2 S-trimer in complex with fab L4.65 and L5.34
Resolution: 2.75 Å
EM Method: Single-particle
Fitted PDBs: 8hp9
Q-score: 0.368
Guo S, Zheng Y, Gao Z, Duan M, Liu S, Du P, Xu X, Xu K, Zhao X, Chai Y, Wang P, Zhao Q, Gao GF, Dai L
Cell Discov (2023) 9 pp. 79-79 [ Pubmed: 37507370 DOI: doi:10.1038/s41421-023-00585-5 ]
- Fab l4.65 (24 kDa, Protein from Homo sapiens)
- Fab l4.65 (23 kDa, Protein from Homo sapiens)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
- Spike protein s2' (125 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Cryo-em structure of sars-cov-2 omicron ba.2 s-trimer in complex with fab l4.65 and l5.34 (Complex from Severe acute respiratory syndrome coronavirus 2)
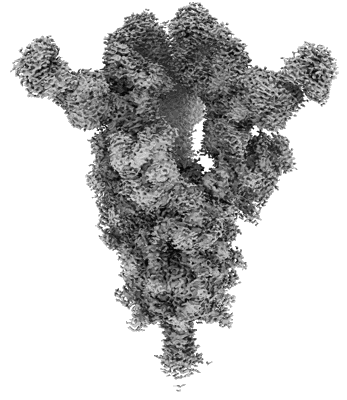
Cryo-EM structure of SARS-CoV-2 Omicron BA.4 S-trimer in complex with fab L4.65 and L5.34
Resolution: 2.85 Å
EM Method: Single-particle
Fitted PDBs: 8hpq
Q-score: 0.243
Guo S, Zheng Y, Gao Z, Duan M, Liu S, Du P, Xu X, Xu K, Zhao X, Chai Y, Wang P, Zhao Q, Gao GF, Dai L
Cell Discov (2023) 9 pp. 79-79 [ Pubmed: 37507370 DOI: doi:10.1038/s41421-023-00585-5 ]
- Fab l4.65 (24 kDa, Protein from Homo sapiens)
- Fab l4.65 (23 kDa, Protein from Homo sapiens)
- Spike protein s2' (125 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
- Cryo-em structure of sars-cov-2 omicron ba.4 s-trimer in complex with fab l4.65 and l5.34 (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Cryo-EM structure of SARS-CoV-2 Omicron Prototype S-trimer in complex with fab L4.65 and L5.34
Resolution: 2.65 Å
EM Method: Single-particle
Fitted PDBs: 8hpv
Q-score: 0.34
Guo S, Zheng Y, Gao Z, Duan M, Liu S, Du P, Xu X, Xu K, Zhao X, Chai Y, Wang P, Zhao Q, Gao GF, Dai L
Cell Discov (2023) 9 pp. 79-79 [ Pubmed: 37507370 DOI: doi:10.1038/s41421-023-00585-5 ]
- Fab l4.65 (24 kDa, Protein from Homo sapiens)
- Spike protein s2' (125 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Fab l4.65 (23 kDa, Protein from Homo sapiens)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
- Cryo-em structure of sars-cov-2 omicron prototype s-trimer in complex with fab l4.65 and l5.34 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Fab l5.34 (23 kDa, Protein from Homo sapiens)
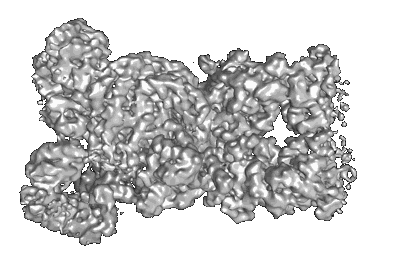
Structure of dimeric mouse SCMC core complex
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 8h93
Q-score: 0.398
Chi P, Ou G, Qin D, Han Z, Li J, Xiao Q, Gao Z, Xu C, Qi Q, Liu Q, Liu S, Li J, Guo L, Lu Y, Chen J, Wang X, Shi H, Li L, Deng D
Nat Struct Mol Biol (2024) 31 pp. 115-124 [ Pubmed: 38177687 DOI: doi:10.1038/s41594-023-01153-x ]
- Oocyte-expressed protein homolog (18 kDa, Protein from Mus musculus)
- Nacht, lrr and pyd domains-containing protein 5 (119 kDa, Protein from Mus musculus)
- Transducin-like enhancer protein 6 (65 kDa, Protein from Mus musculus)
- The mouse subcortical maternal complex (Complex from Mus musculus)