Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
CryoEM structure of dsDNA-RuvB-RuvA domain3 complex
Resolution: 7.02 Å
EM Method: Single-particle
Fitted PDBs: 7x7p
Q-score: 0.186
Zhang X, Zhou Z, Dai L, Chao Y, Liu Z, Huang M, Qu Q, Lin Z
Front Plant Sci (2023) 14 pp. 1139106-1139106 [ Pubmed: 37025142 DOI: doi:10.3389/fpls.2023.1139106 ]
- Dna (7 kDa, DNA from synthetic construct)
- Dna (Complex from Pseudomonas aeruginosa PAO1)
- Ruvb region of the ruva-ruvb-holliday junction complex (Complex from synthetic construct)
- Ruvb-ruva (Complex)
- Holliday junction atp-dependent dna helicase ruva (4 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Holliday junction atp-dependent dna helicase ruvb (34 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Dna (7 kDa, DNA from synthetic construct)
CryoEM structure of RuvA-RuvB-Holliday junction complex
Resolution: 7.02 Å
EM Method: Single-particle
Fitted PDBs: 7x7q
Q-score: 0.167
Zhang X, Zhou Z, Dai L, Chao Y, Liu Z, Huang M, Qu Q, Lin Z
Front Plant Sci (2023) 14 pp. 1139106-1139106 [ Pubmed: 37025142 DOI: doi:10.3389/fpls.2023.1139106 ]
- Ruva-ruvb (Complex from synthetic construct)
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
- Holliday junction atp-dependent dna helicase ruvb (38 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Ruva-ruvb-holliday junction complex (Complex from Pseudomonas aeruginosa PAO1)
- Dna (40-mer) (12 kDa, DNA from synthetic construct)
- Holliday junction atp-dependent dna helicase ruva (21 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Dna (Complex)
- Dna (40-mer) (12 kDa, DNA from synthetic construct)
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
CryoEM structure of RuvA-Holliday junction complex
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 7x5a
Q-score: 0.5
Zhang X, Zhou Z, Dai L, Chao Y, Liu Z, Huang M, Qu Q, Lin Z
Front Plant Sci (2023) 14 pp. 1139106-1139106 [ Pubmed: 37025142 DOI: doi:10.3389/fpls.2023.1139106 ]
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
- Holliday junction (Complex)
- Ruva-holliday junction complex (Complex)
- Holliday junction atp-dependent dna helicase ruva (15 kDa, Protein from Pseudomonas aeruginosa PAO1)
- Ruva (Complex from Pseudomonas aeruginosa PAO1)
- Dna (26-mer) (7 kDa, DNA from synthetic construct)
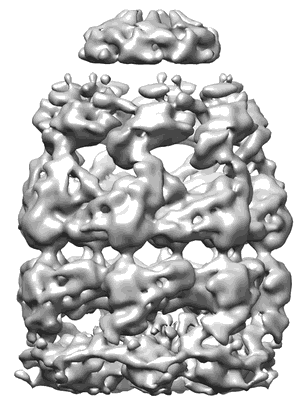
Visualizing GroEL/ES in the Act of Encapsulating a Non-Native Substrate Protein
Resolution: 8.9 Å
EM Method: Single-particle
Fitted PDBs: 3zpz
Q-score: 0.138
Chen D-H, Madan D, Weaver J, Lin Z, Schroder GF, Chiu W, Rye HS
Cell (2013) 153 pp. 1354-1365 [ Pubmed: 23746846 DOI: doi:10.1016/j.cell.2013.04.052 ]
- Groel variant el43py capped by groes with the assistance of nucleotide atp (870 kDa, Sample)
- Groelcys0-d398a-s43c-pyrene (800 kDa, Protein from Escherichia coli K-12)
- Adenosine triphosphate (Ligand from synthetic construct)
- Groes (70 kDa, Protein from Escherichia coli K-12)
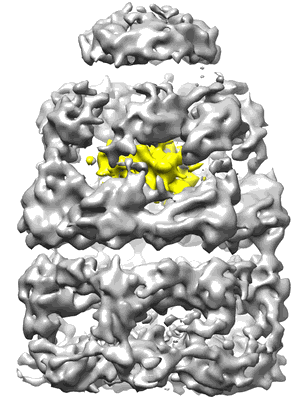
Visualizing GroEL/ES in the Act of Encapsulating a Non-Native Substrate Protein
Resolution: 9.2 Å
EM Method: Single-particle
Fitted PDBs: 3zq0
Q-score: 0.114
Chen D-H, Madan D, Weaver J, Lin Z, Schroder GF, Chiu W, Rye HS
Cell (2013) 153 pp. 1354-1365 [ Pubmed: 23746846 DOI: doi:10.1016/j.cell.2013.04.052 ]
- Groelcys0-d398a-s43c-pyrene (800 kDa, Protein from Escherichia coli K-12)
- Ribulose-1,5-bisphosphate carboxylase oxygenase (50 kDa, Protein from Rhodospirillum rubrum)
- Groes (70 kDa, Protein from Escherichia coli K-12)
- Non-native rubisco substrate protein encapsulated inside the cavity of groel variant el43py capped by groes with the assistance of nucleotide atp (920 kDa, Sample)
- Adenosine triphosphate (Ligand from synthetic construct)
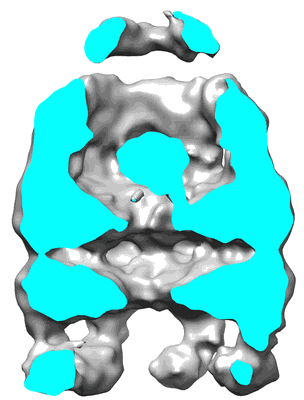
Visualizing GroEL/ES in the Act of Encapsulating a Non-Native Substrate Protein
Resolution: 15.9 Å
EM Method: Single-particle
Fitted PDBs: 3zq1
Q-score: 0.067
Chen D-H, Madan D, Weaver J, Lin Z, Schroder GF, Chiu W, Rye HS
Cell (2013) 153 pp. 1354-1365 [ Pubmed: 23746846 DOI: doi:10.1016/j.cell.2013.04.052 ]
- Groel-d398a (800 kDa, Protein from Escherichia coli K-12)
- Ribulose-1,5-bisphosphate carboxylase oxygenase (50 kDa, Protein from Rhodospirillum rubrum)
- Groes (70 kDa, Protein from Escherichia coli K-12)
- Non-native rubisco substrate protein encapsulated inside the cavity of groel capped by groes with the assistance of nucleotide atp (920 kDa, Sample)
- Adenosine triphosphate (Ligand from synthetic construct)