Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8qma
Q-score: 0.461
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910- [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
- Pap9 (34 kDa, Protein from Sinapis alba)
- Pap3 (79 kDa, Protein from Sinapis alba)
- Pap7 (55 kDa, Protein from Sinapis alba)
- Pap12 (dna-directed rna polymerase subunit omega) (18 kDa, Protein from Sinapis alba)
- Fe (iii) ion (55 Da, Ligand)
- Dna-directed rna polymerase subunit beta'' (156 kDa, Protein from Sinapis alba)
- Pap4 (30 kDa, Protein from Sinapis alba)
- Dna-directed rna polymerase subunit beta' (78 kDa, Protein from Sinapis alba)
- S-adenosylmethionine (398 Da, Ligand)
- Plastid-encoded dna-dependent rna polymerase (pep) (1 MDa, Complex from Sinapis alba)
- Pap11 (85 kDa, Protein from Sinapis alba)
- Dna-directed rna polymerase subunit beta (121 kDa, Protein from Sinapis alba)
- Pap5 (60 kDa, Protein from Sinapis alba)
- Dna-directed rna polymerase subunit alpha (37 kDa, Protein from Sinapis alba)
- Pap1 (103 kDa, Protein from Sinapis alba)
- Pap6 (52 kDa, Protein from Sinapis alba)
- Pap10 (20 kDa, Protein from Sinapis alba)
- Zinc ion (65 Da, Ligand)
- Pap8 (38 kDa, Protein from Sinapis alba)
- Ptac18 (16 kDa, Protein from Sinapis alba)
- Fln2 (68 kDa, Protein from Sinapis alba)
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba - Map A
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910-925.e5 [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba - Map B.
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910-925.e5 [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba - Map D
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910-925.e5 [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba.- Map E
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910-925.e5 [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
Structure of the plastid-encoded RNA polymerase complex (PEP) from Sinapis alba - Map F
do Prado PFV, Ahrens FM, Liebers M, Ditz N, Braun HP, Pfannschmidt T, Hillen HS
Mol Cell (2024) 84 pp. 910-925.e5 [ DOI: doi:10.1016/j.molcel.2024.02.003 Pubmed: 38428434 ]
Structural basis of GTPase-mediated mitochondrial ribosome biogenesis and recycling - dataset2
Methods in molecular biology (Clifton, N.J.) (2023) 2661 pp. 89-100 [ Pubmed: 37166633 DOI: 10.1007/978-1-0716-3171-3_6 ]
Image sets:
Single particle cryo-EM dataset of Sars-CoV2 Rna dependent RNA polymerase
Jochheim FA, Tegunov D, Hillen HS, Schmitzová J, Kokic G, Dienemann C, Cramer P
Commun Biol (2021) 4 [ Pubmed: 34429502 DOI: 10.1038/s42003-021-02529-9 ]
Image sets:
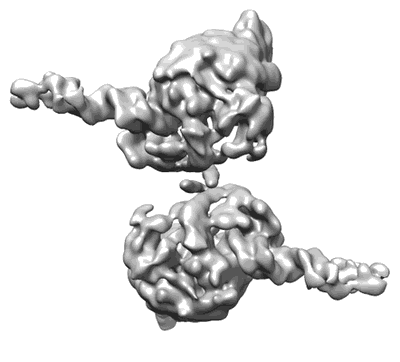
Dimeric form of SARS-CoV-2 RNA-dependent RNA polymerase
Resolution: 5.5 Å
EM Method: Single-particle
Fitted PDBs: 7oyg
Q-score: 0.151
Jochheim FA, Tegunov D, Hillen HS, Schmitzova J, Kokic G, Dienemann C, Cramer P
Communications Biology (2021) 4 pp. 999- [ DOI: doi:10.1038/s42003-021-02529-9 ]
- Sars-cov-2 nsp7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*up*gp*cp*ap*cp*up*gp*cp*gp*up*ap*g)-3') (18 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 rna-dependent rna polymerase (nsp12) (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Dimeric form of sars-cov-2 rna-dependent rna polymerase complex (nsp12, nsp7, nsp8) (Complex from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 nsp8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*up*ap*cp*gp*cp*ap*gp*up*g)-3') (4 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)