Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
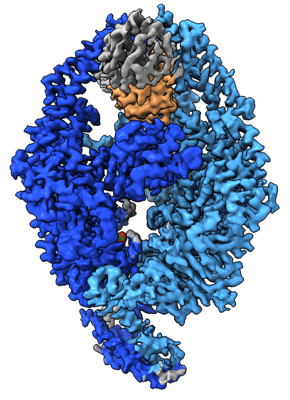
human MutSalpha (MSH2/MSH6) binding to DNA with a GT mismatch
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8ag6
Q-score: 0.542
Bruekner SR, Pieters W, Fish A, Liaci AM, Scheffers S, Rayner E, Kaldenbach D, Drost L, Dekker M, van Hees-Stuivenberg S, Delzenne-Goette E, de Konink C, Houlleberghs H, Dubbink HJ, AlSaegh A, de Wind N, Forster F, Te Riele H, Sixma TK
Nucleic Acids Res (2023) 51 pp. 1173-1188 [ Pubmed: 36715327 DOI: doi:10.1093/nar/gkad015 ]
- Dna mismatch repair protein msh6 (158 kDa, Protein from Homo sapiens)
- Msh2 (Complex from Homo sapiens)
- Dna (50-mer) (15 kDa, DNA from synthetic construct)
- Dna (50-mer) (15 kDa, DNA from synthetic construct)
- Mutsalpha on mismatched dna (31 kDa, Complex from Homo sapiens)
- Msh6 (Complex from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Dna containing a g/t mismatch (Complex from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Dna mismatch repair protein msh2 (104 kDa, Protein from Homo sapiens)
human MutSalpha mutant (MSH2-V63E/MSH6) on DNA containing a GT mismatch
Bruekner SR, Pieters W, Fish A, Liaci AM, Scheffers S, Rayner E, Kaldenbach D, Drost L, Dekker M, van Hees-Stuivenberg S, Delzenne-Goette E, de Konink C, Houllleberghs H, Dubbink HJ, AlSaegh A, de Wind N, Forster F, te Riele H, Sixma TK
Nucleic Acids Res (2023) [ ]
- Human mutsalpha mutant (msh2-v63e/msh6) on mismatched dna (31 kDa, Complex from Homo sapiens)
- Msh6 (Complex from Homo sapiens)
- Dna (50-mer) (DNA from synthetic construct)
- Dna (50-mer) (Complex from synthetic construct)
- Msh6 (Protein from Homo sapiens)
- Msh2 v63e (Protein from Homo sapiens)
- Msh2 v63e (Complex from Homo sapiens)
human MutSalpha (MSH2/MSH6) on DNA containing a GT mismatch in the presence of ADP
Bruekner SR, Pieters W, Fish A, Liaci AM, Scheffers S, Rayner E, Kaldenbach D, Drost L, Dekker M, van Hees-Stuivenberg S, Delzenne-Goette E, de Konink C, Houlleberghs H, Dubbink HJ, AlSaegh A, de Wind N, Forster F, Te Riele H, Sixma TK
Nucleic Acids Res (2023) 51 pp. 1173-1188 [ Pubmed: 36715327 DOI: doi:10.1093/nar/gkad015 ]
- Msh2 (Complex from Homo sapiens)
- Msh2 (Protein from Homo sapiens)
- Msh6 (Complex from Homo sapiens)
- Dna (50-mer) (DNA from synthetic construct)
- Mutsalpha on mismatched dna in the presence of adp (31 kDa, Complex from Homo sapiens)
- Dna (50-mer) (Complex from synthetic construct)
- Msh6 (Protein from Homo sapiens)
Resolution: 4.4 Å
EM Method: Single-particle
Fitted PDBs: 7ai5
Q-score: 0.239
Fernandez-Leiro R, Bhairosing-Kok D, Kunetsky V, Laffeber C, Winterwerp HH, Groothuizen F, Fish A, Lebbink JHG, Friedhoff P, Sixma TK, Lamers MH
Nat Struct Mol Biol (2021) 28 pp. 373-381 [ DOI: doi:10.1038/s41594-021-00577-7 Pubmed: 33820992 ]
- Dna (5'-d(p*cp*gp*gp*tp*ap*cp*cp*cp*ap*ap*tp*tp*cp*gp*cp*cp*cp*tp*ap*tp*ap*g)-3') (6 kDa, DNA from synthetic construct)
- Dna (Complex from synthetic construct)
- Dna mismatch repair protein muts (Complex from Escherichia coli)
- Dna mismatch repair protein muts (95 kDa, Protein from Escherichia coli)
- Muts loaded on matched dna in the presence of atp (190 kDa, Complex)
- Dna (5'-d(p*cp*tp*ap*tp*ap*gp*gp*gp*cp*gp*ap*ap*tp*tp*gp*gp*gp*tp*ap*cp*cp*g)-3') (6 kDa, DNA from synthetic construct)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Resolution: 6.9 Å
EM Method: Single-particle
Fitted PDBs: 7ai6
Q-score: 0.144
Fernandez-Leiro R, Bhairosing-Kok D, Kunetsky V, Laffeber C, Winterwerp HH, Groothuizen F, Fish A, Lebbink JHG, Friedhoff P, Sixma TK, Lamers MH
Nat Struct Mol Biol (2021) 28 pp. 373-381 [ DOI: doi:10.1038/s41594-021-00577-7 Pubmed: 33820992 ]
- Dna (25-mer) (7 kDa, DNA from synthetic construct)
- Dna mismatch repair protein muts (95 kDa, Protein from Escherichia coli (strain K12))
- Dna (Complex from synthetic construct)
- Dna (25-mer) (7 kDa, DNA from synthetic construct)
- Dna mismatch repair protein muts (Complex from Escherichia coli (strain K12))
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Muts loaded on dna substrate with one central mismatched basepair in the presence of amp-pnp (190 kDa, Complex)
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7ai7
Q-score: 0.336
Fernandez-Leiro R, Bhairosing-Kok D, Kunetsky V, Laffeber C, Winterwerp HH, Groothuizen F, Fish A, Lebbink JHG, Friedhoff P, Sixma TK, Lamers MH
Nat Struct Mol Biol (2021) 28 pp. 373-381 [ DOI: doi:10.1038/s41594-021-00577-7 Pubmed: 33820992 ]
- Dna (5'-d(p*cp*tp*tp*ap*gp*cp*tp*tp*ap*gp*gp*ap*tp*c)-3') (4 kDa, DNA from synthetic construct)
- Dna (Complex from synthetic construct)
- Dna mismatch repair protein muts (Complex from Escherichia coli (strain K12))
- Muts loaded on matched dna in the presence of atp (190 kDa, Complex)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Dna (5'-d(p*gp*ap*tp*cp*cp*tp*ap*ap*cp*tp*ap*ap*g)-3') (4 kDa, DNA from synthetic construct)
- Dna mismatch repair protein muts (95 kDa, Protein from Escherichia coli (strain K12))
Resolution: 4.7 Å
EM Method: Single-particle
Fitted PDBs: 7aib
Q-score: 0.198
Fernandez-Leiro R, Bhairosing-Kok D, Kunetsky V, Laffeber C, Winterwerp HH, Groothuizen F, Fish A, Lebbink JHG, Friedhoff P, Sixma TK, Lamers MH
Nat Struct Mol Biol (2021) 28 pp. 373-381 [ DOI: doi:10.1038/s41594-021-00577-7 Pubmed: 33820992 ]
- Muts loaded on dna substrate with one central mismatched basepair in the presence of amp-pnp (280 kDa, Complex)
- Dna (30-mer) (9 kDa, DNA from synthetic construct)
- Dna mismatch repair protein muts (95 kDa, Protein from Escherichia coli (strain K12))
- Dna (Complex from synthetic construct)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Dna mismatch repair protein muts and dna mismatch repair endonuclease mutl (Complex from Escherichia coli (strain K12))
- Dna mismatch repair protein mutl (39 kDa, Protein from Escherichia coli (strain K12))
- Dna (30-mer) (9 kDa, DNA from synthetic construct)
MutS-MutL in clamp state (kinked clamp domain)
Resolution: 5.0 Å
EM Method: Single-particle
Fitted PDBs: 7aic
Q-score: 0.158
Fernandez-Leiro R, Bhairosing-Kok D, Kunetsky V, Laffeber C, Winterwerp HH, Groothuizen F, Fish A, Lebbink JHG, Friedhoff P, Sixma TK, Lamers MH
Nat Struct Mol Biol (2021) 28 pp. 373-381 [ DOI: doi:10.1038/s41594-021-00577-7 Pubmed: 33820992 ]
- Muts loaded on dna substrate with one central mismatched basepair in the presence of amp-pnp (280 kDa, Complex)
- Dna (30-mer) (9 kDa, DNA from synthetic construct)
- Dna mismatch repair protein muts (95 kDa, Protein from Escherichia coli (strain K12))
- Dna (Complex from synthetic construct)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Dna mismatch repair protein muts and dna mismatch repair endonuclease mutl (Complex from Escherichia coli (strain K12))
- Dna mismatch repair protein mutl (39 kDa, Protein from Escherichia coli (strain K12))
- Dna (30-mer) (9 kDa, DNA from synthetic construct)