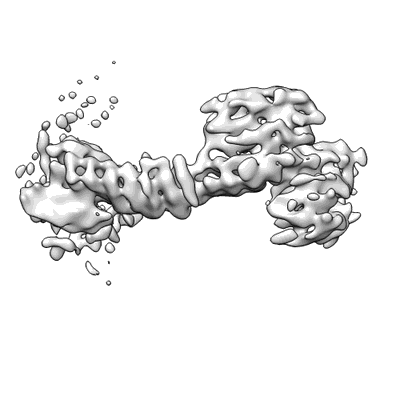Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
In situ structure of HDCR filaments
Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Trischler R, Schuller SK, Wagner J, Schwarz FM, Engel BD, Muller V, Schuller JM
Nature (2022) 607 pp. 823-830 [ Pubmed: 35859174 DOI: doi:10.1038/s41586-022-04971-z ]
In situ structure of the T. kivui ribosome
Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Trischler R, Schuller SK, Wagner J, Schwarz FM, Engel BD, Muller V, Schuller JM
Nature (2022) 607 pp. 823-830 [ Pubmed: 35859174 DOI: doi:10.1038/s41586-022-04971-z ]
In situ cryo-electron tomogram of a T. kivui cell 1
Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Trischler R, Schuller SK, Wagner J, Schwarz FM, Engel BD, Muller V, Schuller JM
Nature (2022) 607 pp. 823-830 [ Pubmed: 35859174 DOI: doi:10.1038/s41586-022-04971-z ]
In situ cryo-electron tomogram of a T. kivui cell 2
Dietrich HM, Righetto RD, Kumar A, Wietrzynski W, Trischler R, Schuller SK, Wagner J, Schwarz FM, Engel BD, Muller V, Schuller JM
Nature (2022) 607 pp. 823-830 [ Pubmed: 35859174 DOI: doi:10.1038/s41586-022-04971-z ]
Resolution: 2.13 Å
EM Method: Single-particle
Fitted PDBs: 7jfo
Q-score: 0.734
He S, Chou HT, Matthies D, Wunder T, Meyer MT, Atkinson N, Martinez-Sanchez A, Jeffrey PD, Port SA, Patena W, He G, Chen VK, Hughson FM, McCormick AJ, Mueller-Cajar O, Engel BD, Yu Z, Jonikas MC
Nat Plants (2020) 6 pp. 1480-1490 [ DOI: doi:10.1038/s41477-020-00811-y Pubmed: 33230314 ]
- Ribulose bisphosphate carboxylase small chain 2, chloroplastic (20 kDa, Protein from Chlamydomonas reinhardtii)
- Epyc1(49-72) peptide-bound rubisco (Complex from Chlamydomonas reinhardtii)
- Lci5 (2 kDa, Protein from Chlamydomonas reinhardtii)
- Ribulose bisphosphate carboxylase large chain (52 kDa, Protein from Chlamydomonas reinhardtii)
- Water (18 Da, Ligand)
Resolution: 2.68 Å
EM Method: Single-particle
Fitted PDBs: 7jn4
Q-score: 0.624
He S, Chou HT, Matthies D, Wunder T, Meyer MT, Atkinson N, Martinez-Sanchez A, Jeffrey PD, Port SA, Patena W, He G, Chen VK, Hughson FM, McCormick AJ, Mueller-Cajar O, Engel BD, Yu Z, Jonikas MC
Nat Plants (2020) 6 pp. 1480-1490 [ DOI: doi:10.1038/s41477-020-00811-y Pubmed: 33230314 ]
- Rubisco at apo state (Complex from Chlamydomonas reinhardtii)
- Ribulose bisphosphate carboxylase large chain (52 kDa, Protein from Chlamydomonas reinhardtii)
- Water (18 Da, Ligand)
- Ribulose bisphosphate carboxylase small chain 2, chloroplastic (20 kDa, Protein from Chlamydomonas reinhardtii)
EPYC1(106-135) peptide-bound Rubisco
Resolution: 2.06 Å
EM Method: Single-particle
Fitted PDBs: 7jsx
Q-score: 0.677
He S, Chou HT, Matthies D, Wunder T, Meyer MT, Atkinson N, Martinez-Sanchez A, Jeffrey PD, Port SA, Patena W, He G, Chen VK, Hughson FM, McCormick AJ, Mueller-Cajar O, Engel BD, Yu Z, Jonikas MC
Nat Plants (2020) 6 pp. 1480-1490 [ DOI: doi:10.1038/s41477-020-00811-y Pubmed: 33230314 ]
- Ribulose bisphosphate carboxylase small chain 2, chloroplastic (20 kDa, Protein from Chlamydomonas reinhardtii)
- Epyc1 (3 kDa, Protein from Chlamydomonas reinhardtii)
- Ribulose bisphosphate carboxylase large chain (52 kDa, Protein from Chlamydomonas reinhardtii)
- Water (18 Da, Ligand)
- Epyc1(106-135) peptide-bound rubisco (Complex from Chlamydomonas reinhardtii)
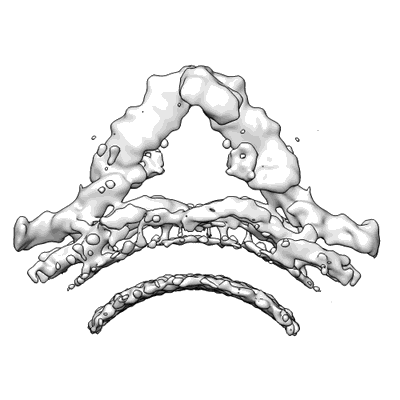
Retromer-Vps5: low-resolution overview map centred on the Vps35/29 arch.
Kovtun O, Leneva N, Bykov YS, Ariotti N, Teasdale RD, Schaffer M, Engel BD, Owen DJ, Briggs JAG, Collins BM
Nature (2018) 561 pp. 561-564 [ Pubmed: 30224749 DOI: doi:10.1038/s41586-018-0526-z ]
- Vacuolar protein sorting-associated protein 29 (Protein)
- Vacuolar protein sorting-associated protein 26 (Protein)
- Retromer-vps5 complex assembled on the membrane. (Complex from Chaetomium thermophilum var. thermophilum DSM 1495)
- Vacuolar protein sorting-associated protein 5 (Protein)
- Vacuolar protein sorting-associated protein 35 (Protein)
Retromer-Vps5: map centred on the Vps35/29 arch.
Kovtun O, Leneva N, Bykov YS, Ariotti N, Teasdale RD, Schaffer M, Engel BD, Owen DJ, Briggs JAG, Collins BM
Nature (2018) 561 pp. 561-564 [ Pubmed: 30224749 DOI: doi:10.1038/s41586-018-0526-z ]
Retromer-Vps5: map centred on the leg of the Vps35/29 arch.
Kovtun O, Leneva N, Bykov YS, Ariotti N, Teasdale RD, Schaffer M, Engel BD, Owen DJ, Briggs JAG, Collins BM
Nature (2018) 561 pp. 561-564 [ Pubmed: 30224749 DOI: doi:10.1038/s41586-018-0526-z ]