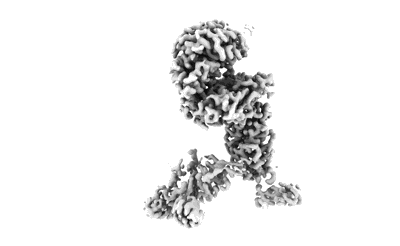Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
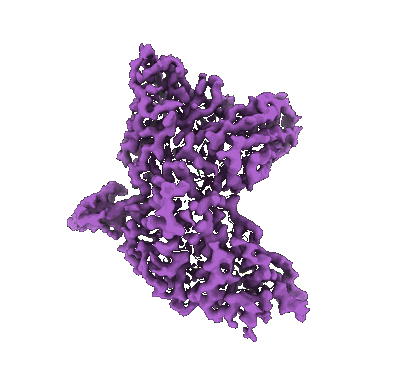
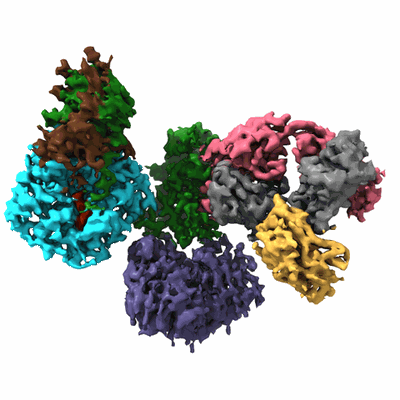
SARS-CoV-2 Omicron Variant S Trimer complexed with two JMB2002 Fab
Resolution: 2.69 Å
EM Method: Single-particle
Fitted PDBs: 7wpe
Q-score: 0.338
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Sars-cov-2 omicron variant s trimer complexed with jmb2002 fab (Complex from Severe acute respiratory syndrome coronavirus)
- Anti-fab nanobody (Complex)
- Sars-cov-2 omicron variant s (Complex from Mus musculus)
- Jmb2002 fab heavy chain (25 kDa, Protein from Mus musculus)
- Jmb2002 fab light chian (23 kDa, Protein from Mus musculus)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Jmb2002 fab (Complex from Lama glama)
- Anti-fab nanobody (14 kDa, Protein from Lama glama)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
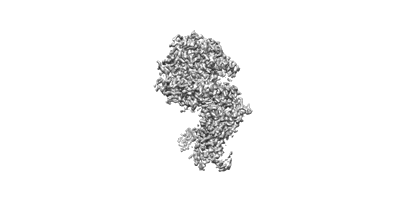
The interface of JMB2002 Fab binds to SARS-CoV-2 Omicron Variant S
Resolution: 2.47 Å
EM Method: Single-particle
Fitted PDBs: 7wrv
Q-score: 0.507
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Sars-cov-2 omicron variant s (Complex from Mus musculus)
- Jmb2002 fab (Complex)
- Jmb2002 fab heavy chain (25 kDa, Protein from Mus musculus)
- Jmb2002 fab light chain (23 kDa, Protein from Mus musculus)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron variant s trimer complexed with jmb2002 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 Omicron Variant S Trimer complexed with one JMB2002 Fab
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 7wpd
Q-score: 0.265
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Anti fab nanobody (Complex)
- Jmb2002 fab heavy chain (25 kDa, Protein from Mus musculus)
- Jmb2002 fab light chian (23 kDa, Protein from Mus musculus)
- Sars-cov-2 omicron variant s trimer (Complex from Mus musculus)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Jmb2002 fab (Complex from Lama glama)
- Sars-cov-2 omicron variant s trimer complexed with jmb2002 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
- Anti-fab nanobody (14 kDa, Protein from Lama glama)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
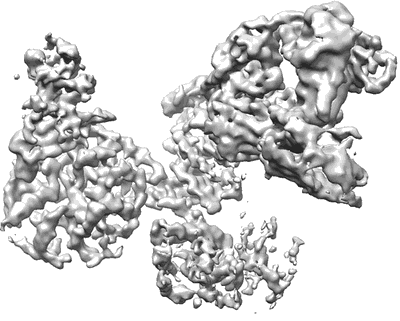
SARS-CoV-2 Omicron Variant S Trimer complexed with three JMB2002 Fab
Resolution: 2.92 Å
EM Method: Single-particle
Fitted PDBs: 7wpf
Q-score: 0.226
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 omicron variant s (Complex from Mus musculus)
- Jmb2002 fab heavy chain (25 kDa, Protein from Mus musculus)
- Anti fab nanobody (Complex)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Jmb2002 fab (Complex from Lama glama)
- Sars-cov-2 omicron variant s trimer complexed with jmb2002 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
- Anti-fab nanobody (14 kDa, Protein from Lama glama)
- Jmb2002 fab light chain (23 kDa, Protein from Mus musculus)
SARS-CoV-2 Omicron Variant SPIKE trimer, all RBDs down
Resolution: 2.56 Å
EM Method: Single-particle
Fitted PDBs: 7wp9
Q-score: 0.377
SARS-CoV-2 Omicron Variant SPIKE trimer complexed with ACE2
Resolution: 2.77 Å
EM Method: Single-particle
Fitted PDBs: 7wpa
Q-score: 0.371
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Chloride ion (35 Da, Ligand)
- Sars-cov-2 omicron variant spike trimer complexed with ace2 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Angiotensin-converting enzyme 2 (92 kDa, Protein from Homo sapiens)
- Sars-cov-2 omicron variant spike (Complex from Homo sapiens)
- Ace2 (Complex)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 Omicron Variant RBD complexed with ACE2
Resolution: 2.79 Å
EM Method: Single-particle
Fitted PDBs: 7wpb
Q-score: 0.599
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Chloride ion (35 Da, Ligand)
- Sars-cov-2 omicron variant rbd (Complex from Homo sapiens)
- Angiotensin-converting enzyme 2 (92 kDa, Protein from Homo sapiens)
- Ace2 (Complex)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 omicron variant rbd complexed with ace2 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
The second RBD of SARS-CoV-2 Omicron Variant in complexed with RBD-ACE2
Resolution: 2.57 Å
EM Method: Single-particle
Fitted PDBs: 7wpc
Q-score: 0.405
Yin W, Xu Y, Xu P, Cao X, Wu C, Gu C, He X, Wang X, Huang S, Yuan Q, Wu K, Hu W, Huang Z, Liu J, Wang Z, Jia F, Xia K, Liu P, Wang X, Song B, Zheng J, Jiang H, Cheng X, Jiang Y, Deng SJ, Xu HE
Science (2022) 375 pp. 1048-1053 [ Pubmed: 35133176 DOI: doi:10.1126/science.abn8863 ]
- Chloride ion (35 Da, Ligand)
- Sars-cov-2 omicron variant rbd (Complex from Homo sapiens)
- Angiotensin-converting enzyme 2 (92 kDa, Protein from Homo sapiens)
- The second rbd of sars-cov-2 omicron variant in complexed with rbd-ace2 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Ace2 (Complex)
- Spike glycoprotein (133 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
ATP-bound AMP-activated protein kinase
Resolution: 3.93 Å
EM Method: Single-particle
Fitted PDBs: 7m74
Q-score: 0.27
Yan Y, Mukherjee S, Harikumar KG, Strutzenberg TS, Zhou XE, Suino-Powell K, Xu TH, Sheldon RD, Lamp J, Brunzelle JS, Radziwon K, Ellis A, Novick SJ, Vega IE, Jones RG, Miller LJ, Xu HE, Griffin PR, Kossiakoff AA, Melcher K
Science (2021) 373 pp. 413-419 [ Pubmed: 34437114 DOI: doi:10.1126/science.abe7565 ]
- 5'-amp-activated protein kinase catalytic subunit alpha-1 (56 kDa, Protein from Homo sapiens)
- 5'-amp-activated protein kinase subunit beta-2 (22 kDa, Protein from Homo sapiens)
- Maltose/maltodextrin abc transporter substrate-binding protein male (40 kDa, Protein from Escherichia coli)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Mbp-fused atp bound ampk in complex with c-compound stabilized by fab and a nanobody (Complex from Homo sapiens)
- Nanobody (13 kDa, Protein from Lama glama)
- 5'-amp-activated protein kinase subunit gamma-1 (34 kDa, Protein from Homo sapiens)
- Fab light chain (22 kDa, Protein from Homo sapiens)
- 6-[4-(2-piperidin-1-ylethoxy)phenyl]-3-pyridin-4-ylpyrazolo[1,5-a]pyrimidine (399 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Adenosine monophosphate (347 Da, Ligand)
- Fab heavy chain (24 kDa, Protein from Homo sapiens)
Resolution: 3.47 Å
EM Method: Single-particle
Fitted PDBs: 7jhg
Q-score: 0.345
Yan Y, Mukherjee S, Harikumar KG, Strutzenberg TS, Zhou XE, Suino-Powell K, Xu TH, Sheldon RD, Lamp J, Brunzelle JS, Radziwon K, Ellis A, Novick SJ, Vega IE, Jones RG, Miller LJ, Xu HE, Griffin PR, Kossiakoff AA, Melcher K
Science (2021) 373 pp. 413-419 [ Pubmed: 34437114 DOI: doi:10.1126/science.abe7565 ]
- Fab heavy chain (25 kDa, Protein from synthetic construct)
- 5'-amp-activated protein kinase catalytic subunit alpha-1 (56 kDa, Protein from Homo sapiens)
- 5'-amp-activated protein kinase subunit beta-2 (22 kDa, Protein from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Maltodextrin-binding protein (40 kDa, Protein from Escherichia coli K-12)
- 5'-amp-activated protein kinase subunit gamma-1 (34 kDa, Protein from Homo sapiens)
- Cryo-em structure of atp-bound fully inactive ampk in complex with dorsomorphin (compound c) and fab-nanobody (220 kDa, Complex)
- Fab light chain (23 kDa, Protein from synthetic construct)
- 6-[4-(2-piperidin-1-ylethoxy)phenyl]-3-pyridin-4-ylpyrazolo[1,5-a]pyrimidine (399 Da, Ligand)
- Fab-nanobody (Complex from synthetic construct)
- Nanobody (17 kDa, Protein from synthetic construct)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Adenosine monophosphate (347 Da, Ligand)
- Atp-bound fully inactive ampk in complex with dorsomorphin (compound c) (Complex from Homo sapiens)