 |
PDBsum entry 2jzf

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Viral protein
|
PDB id
|
|

|

|
2jzf
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
* Residue conservation analysis
|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Viral protein
|
 |
|
Title:
|
 |
Nmr conformer closest to the mean coordinates of the domain 513-651 of the sars-cov nonstructural protein nsp3
|
|
Structure:
|
 |
Replicase polyprotein 1ab. Chain: a. Fragment: non-structural protein 3 (domain 513-651): residues 1331- 1469. Synonym: pp1ab, orf1ab polyprotein [includes: replicase polyprotein 1a, non-structural proteins 1,2,3,4, 3c-like proteinase, non- structural proteins 6,7,8,9,10, RNA-directed RNA polymerase, helicase, exoribonuclease, uridylate-specific endoribonuclease, putative 2'-o-methyl transferase].
|
|
Source:
|
 |
Sars coronavirus. Organism_taxid: 227859. Gene: rep, 1a-1b. Expressed in: escherichia coli bl21(de3). Expression_system_taxid: 469008.
|
|
NMR struc:
|
 |
1 models
|
 |
|
Authors:
|
 |
A.Chatterjee,M.A.Johnson,P.Serrano,B.Pedrini,J.Joseph,K.Saikatendu, B.Neuman,R.C.Stevens,I.A.Wilson,M.J.Buchmeier,P.Kuhn,K.Wuthrich, Joint Center For Structural Genomics (Jcsg)
|
|
Key ref:
|
 |
A.Chatterjee
et al.
(2009).
Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
J Virol,
83,
1823-1836.
PubMed id:
|
 |
|
Date:
|
 |
|
04-Jan-08
|
Release date:
|
05-Feb-08
|
|
|
Supersedes:
|

|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P0C6X7
(R1AB_CVHSA) -
Replicase polyprotein 1ab from Severe acute respiratory syndrome coronavirus
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
7073 a.a.
143 a.a.*
|

|

|

|

|
|

|

|

|
 |
 |

|
|
Key: |
 |
PfamA domain |
 |
 |
 |
Secondary structure |
 |
 |
CATH domain |
 |
|
*
PDB and UniProt seqs differ
at 4 residue positions (black
crosses)
|
|

|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.1.1.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
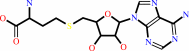 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
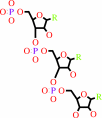 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
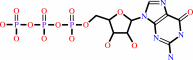 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.1.13.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 8:
|
 |
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 9:
|
 |
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 10:
|
 |
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
Enzyme class 11:
|
 |
E.C.3.6.4.12
- Dna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 12:
|
 |
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 13:
|
 |
E.C.4.6.1.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
J Virol
83:1823-1836
(2009)
|
|
PubMed id:
|
|
|
|

|
| |
|
Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
|
|
A.Chatterjee,
M.A.Johnson,
P.Serrano,
B.Pedrini,
J.S.Joseph,
B.W.Neuman,
K.Saikatendu,
M.J.Buchmeier,
P.Kuhn,
K.Wüthrich.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The nuclear magnetic resonance (NMR) structure of a central segment of the
previously annotated severe acute respiratory syndrome (SARS)-unique domain
(SUD-M, for "middle of the SARS-unique domain") in SARS coronavirus (SARS-CoV)
nonstructural protein 3 (nsp3) has been determined. SUD-M(513-651) exhibits a
macrodomain fold containing the nsp3 residues 528 to 648, and there is a
flexibly extended N-terminal tail with the residues 513 to 527 and a C-terminal
flexible tail of residues 649 to 651. As a follow-up to this initial result, we
also solved the structure of a construct representing only the globular domain
of residues 527 to 651 [SUD-M(527-651)]. NMR chemical shift perturbation
experiments showed that SUD-M(527-651) binds single-stranded poly(A) and
identified the contact area with this RNA on the protein surface, and
electrophoretic mobility shift assays then confirmed that SUD-M has higher
affinity for purine bases than for pyrimidine bases. In a further search for
clues to the function, we found that SUD-M(527-651) has the closest
three-dimensional structure homology with another domain of nsp3, the
ADP-ribose-1"-phosphatase nsp3b, although the two proteins share only 5%
sequence identity in the homologous sequence regions. SUD-M(527-651) also shows
three-dimensional structure homology with several helicases and nucleoside
triphosphate-binding proteins, but it does not contain the motifs of catalytic
residues found in these structural homologues. The combined results from NMR
screening of potential substrates and the structure-based homology studies now
form a basis for more focused investigations on the role of the SARS-unique
domain in viral infection.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
K.R.Hurst,
R.Ye,
S.J.Goebel,
P.Jayaraman,
and
P.S.Masters
(2010).
An interaction between the nucleocapsid protein and a component of the replicase-transcriptase complex is crucial for the infectivity of coronavirus genomic RNA.
|
| |
J Virol,
84,
10276-10288.
|
 |
|
|
|
|
 |
J.A.Wojdyla,
I.Manolaridis,
E.J.Snijder,
A.E.Gorbalenya,
B.Coutard,
Y.Piotrowski,
R.Hilgenfeld,
and
P.A.Tucker
(2009).
Structure of the X (ADRP) domain of nsp3 from feline coronavirus.
|
| |
Acta Crystallogr D Biol Crystallogr,
65,
1292-1300.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
J.Tan,
C.Vonrhein,
O.S.Smart,
G.Bricogne,
M.Bollati,
Y.Kusov,
G.Hansen,
J.R.Mesters,
C.L.Schmidt,
and
R.Hilgenfeld
(2009).
The SARS-Unique Domain (SUD) of SARS Coronavirus Contains Two Macrodomains That Bind G-Quadruplexes.
|
| |
PLoS Pathog,
5,
e1000428.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
P.Serrano,
M.A.Johnson,
A.Chatterjee,
B.W.Neuman,
J.S.Joseph,
M.J.Buchmeier,
P.Kuhn,
and
K.Wüthrich
(2009).
Nuclear magnetic resonance structure of the nucleic acid-binding domain of severe acute respiratory syndrome coronavirus nonstructural protein 3.
|
| |
J Virol,
83,
12998-13008.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.Perlman,
and
J.Netland
(2009).
Coronaviruses post-SARS: update on replication and pathogenesis.
|
| |
Nat Rev Microbiol,
7,
439-450.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

