It’s the 1950s; the cold war is at its height, rock and roll has swept the charts, and science is advancing greatly, leading to the determination of the 1st protein structures, one of which is featured on October of our 2019 calendar.
The structural study of myoglobin started in parallel to that of haemoglobin. Max Perutz was working on haemoglobin in Cambridge when John Kendrew and his team, also in Cambridge, decided to focus on a molecule a quarter the size of haemoglobin. This smaller size made myoglobin a simpler molecule for the first attempt to establish the molecular structure of a protein. The first crystals obtained using horse heart myoglobin were too small for X-ray analysis at the time. This made Kendrew reconsider the source of the sample and he realised that diving animals store more oxygen in their muscles, therefore have more myoglobin. He purified myoglobin from several creatures (including penguins!) but found that using a chunk of sperm whale muscle gave large crystals with clean X-ray diffraction patterns.
But just getting good crystals was not the only hurdle to solving the structure of myoglobin. Kendrew also had to make the diffraction data interpretable. Solving the phase problem by attaching heavy metals to five distinct sites of the molecule allowed him to begin interpreting his 3D structure data. The resolution for data collection was set to 6 Å which was sufficient to see the overall layout of the molecule. The polypeptide chain was visible as a rod of high electron density, folded in a complex pattern. The tertiary structure revealed a remarkable but unexpected feature of the molecule – its lack of symmetry. In further experiments the resolution was increased to 2 Å, which helped in establishing the secondary structure, with α-helices seen for the first time. The resolution improvement meant Kendrew had to deal with tens of thousands of data points instead of a few hundred, and he pioneered using early computers to manage and analyse those data.
Thanks to these further experiments, Kendrew and his team were also able to determine that the configuration of helices is right-handed and that the molecule consists of a number of segments (helices) separated by irregular regions. At this point of these studies there had been no serious attempt made to identify the side-chains and the primary structure of the myoglobin still remained unknown. Protein sequencing was still in its early days too. Kendrew and his team didn’t stop until they managed to define the exact sequence of myoglobin and they realized that knowledge of the protein sequence helps with the interpretation of the electron density maps.
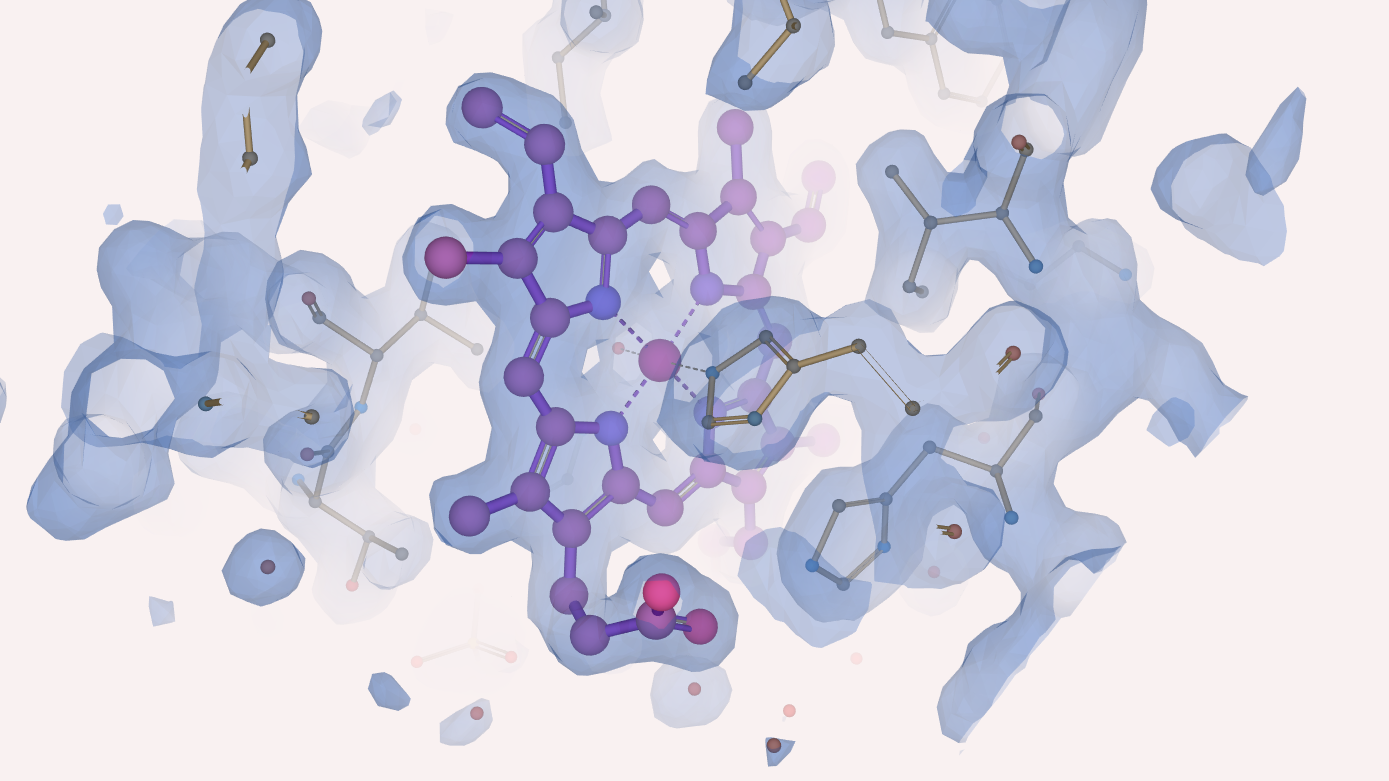
It is well known that myoglobin binds and stores oxygen thanks to presence of the haem group. But how did Kendrew and his team see this porphyrin ring in their experiments and what did they learn about it? With a resolution of 6 Å, the haem was just a disk of high electron density. When they moved to a resolution of 2 Å the iron atom was resolved from the nitrogens of the porphyrin ring and the new results confirmed the orientation of the haem group within the myoglobin, which had been misinterpreted in earlier models. Moreover, they confirmed chemical studies of myoglobin suggesting that the haem group is attached to the protein by a bond from the iron atom to a nitrogen atom of a histidine residue.
In 1962, John Kendrew was awarded the Nobel Prize in Chemistry together with Max Perutz for their studies of the structures of globular proteins (myoglobin and haemoglobin). Myoglobin was one of the first protein structures to be placed in the PDB when it begun in 1971. The 2 Å model of myoglobin (PDB entry 1mbn) was added to the PDB in May 1976. Since then myoglobin has been extensively studied structurally, and there are currently well over 400 structures from 15 different species in the PDB. Despite being one of the most studied proteins, its physiological function is not yet conclusively established.
About the artwork.
Henry from Impington Village College painted this image of oxygenated myoglobin (PDB entry 1mbo) inspired by the artist Gregory Edwards
Romana Gáborová


