EMPIAR data re-use case studies
EMPIAR-10930
Last updated 03/09/2025 by Simone Weyand at EMBL-EBIEMPIAR-10930 showcases how availability of raw cryo-EM data through EMPIAR, combined with advances in processing techniques provide new biological insight
Dataset
Entry EMPIAR-10930 contains the raw cryo-EM data corresponding to the complex of the extracellular ligand binding region (residues 648 – 1025) of anaplastic lymphoma kinase (ALKTG-EGFL) with its ligand ALKAL2 (AUG-alpha) at 2.27 Å resolution, as deposited by the groups of J Schlessinger and CG Kalodimos (Reshetnyak et al., Nature, 2021):
- 2.27 Å resolution cryo-EM structure of anaplastic lymphoma kinase (ALK) extracellular fragment of ligand binding region 648-1025 in complex with AUG-alpha. Reshetnyak AV, Rossi P, Myasnikov AG, Sowaileh M, Mohanty J, Nourse A, Miller DJ, Lax I, Schlessinger J & Kalodimos CG, Nature 600 153-157 (2021).
- PMID: 34819673
- EMDB entry EMD-24095 (image below, left)
- PDB entry 7n00 (image below, right)

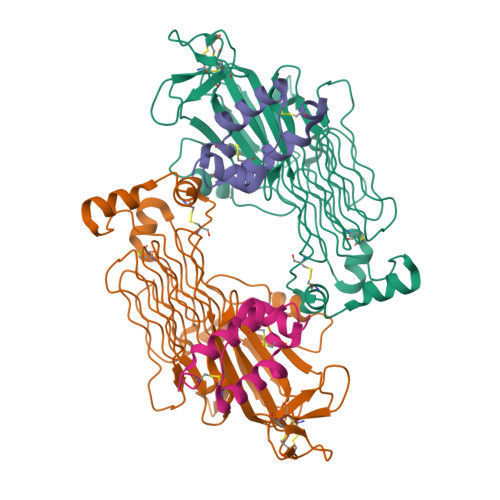
At present, the publication was cited 56 times. The PDB structure has been cited in 9 publications, the EMDB map in 2 publications and the EMPIAR dataset in 1 publication. The dataset is 10.2 TB in size.
The cryo-EM map determined by Reshetnyak et al. corresponds to an ALKTG-EGFL-ALKAL2 complex with 2:2 stoichiometry (Reshetnyak et al., Nature, 2021). On the other hand, the Savvides Lab had determined X-ray structures of the ALK ligand binding region without the membrane-proximal EGFL domain (ALKTG), resulting in a distinct 2:1 stoichiometry for the ALKTG-ALKAL2 complex (De Munck et al., Nature, 2021). While one ALK-ALKAL2 interaction interface is common between the reported 2:1 and 2:2 stoichiometric assemblies, the 2:1 ALKTG-ALKAL2 complex contains an additional ALK receptor interface on ALKAL2 and different ALK receptor-receptor contacts when compared to the 2:2 ALKTG-EGFL-ALKAL2 complex.
Re-processing
By making the raw data publicly available in EMPIAR, other cryo-EM groups have been able to download, analyse and re-process the data. A reanalysis of the data deposited in EMPIAR-10930 by the Savvides group (Felix et al., PLoS Biology, 2025), using the deposited uncleaned particle stack and following a similar initial processing pipeline but without binning the data, revealed the presence of 2D classes corresponding to particles resembling a 2:1 ALKTG-EGFL-ALKAL2 stoichiometry. These particles present roughly 53% of the dataset but suffer severely from preferred orientations that resulted in cryo-EM maps displaying strong anisotropy.
By combining extensive particle orientation rebalancing in cryoSPARC (Punjani et al., Nature Methods, 2017) followed by 3D refinement with Blush regularization in RELION5 (Kimanius et al., Nature Methods, 2024), Felix and colleagues could overcome the map artifacts relating to preferred particle orientations, and report a 3D reconstruction of the 2:1 ALKTG-EGFL-ALKAL2 complex to 3.2 Å resolution from the EMPIAR-10930 dataset (Felix et al., PLoS Biology, 2025). The resulting structure agrees closely with the ALKTG-ALKAL2 X-ray structure with 2:1 stoichiometry reported by De Munck et al., noting that the latter does not contain the membrane-proximal EGFL domains. Therefore, the new ALKTG-EGFL-ALKAL2 complex to 3.2 Å resolution is the most complete complex with 2:1 stoichiometry.
- 3.2 Å resolution cryo-EM structure of a 2:1 ALK:ALKAL2 complex obtained after re- processing of EMPIAR-10930 data. Felix J, De Munck S, Bazan JF, Savvides SN, PLoS Biology 23(4), e3003124 (2025).
- PMID: 40208865
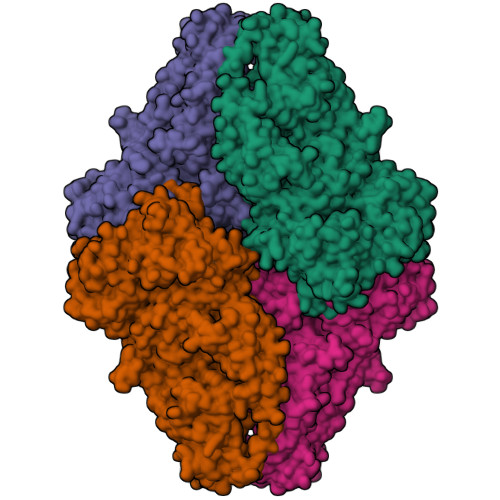
- EMDB entry EMD-51087 (image below, left)
- PDB entry 9g5i (image below, right)


While the biological role of distinct 2:1 and 2:2 ALK-ALKAL2 assemblies in ALK signalling is currently unclear, this analysis provides direct evidence for the presence of an ALKTG-EGFL-ALKAL2 complex with a distinct 2:1 stoichiometry next to the reported 2:2 stoichiometric assembly in the EMPIAR-10930 dataset, and emphasizes the importance of public deposition of raw cryo-EM data to allow reanalysis and interpretation.
References
- Reshetnyak AV, Rossi P, Myasnikov AG, Sowaileh M, Mohanty J, Nourse A, Miller DJ, Lax I, Schlessinger J & Kalodimos CG, Nature 600, 153-157 (2021).
- De Munck S, Provost M, Kurikawa M, Omori I, Mukohyama J, Felix J, Bloch Y, Abdel-Wahab O, Bazan JF, Yoshimi A & Savvides SN, Nature 600, 143-147 (2021).
- Punjani A., Rubinstein JL, Fleet DJ & Brubaker MA. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nature Methods 14, 290-296 (2017).
- Kimanius D, Jamali K, Wilkinsin ME, Lövestam S, Velazhahan V, Nakane T & Scheres SHW. Nature Methods 21, 1216-1221 (2024)
- Felix J, De Munck S, Bazan JF & Savvides SN, PLoS Biology 23(4), e3003124 (2025).
EMPIAR-10061
Last updated 19/02/2020 by Gerard Kleywegt, Ardan Patwardhan & Andrii Iudin at EMBL-EBIEntry EMPIAR-10061 provides a good example of why archiving raw cryo-EM (and general bioimaging) data is important as it enables new science and facilitates and accelerates methods development in this rapidly evolving field.
Dataset
Dataset Entry EMPIAR-10061 (https://empiar.org/10061) contains the raw cryo-EM data underpinning what was a breakthrough structure and at the time the highest resolution cryo-EM structure available, the 2.2Å resolution structure of β-galactosidase by the group of Sriram Subramaniam:
- 2.2 Å resolution cryo-EM structure of β-galactosidase in complex with a cell-permeant inhibitor. Bartesaghi A, Merk A, Banerjee S, Matthies D, Wu X, Milne JL, Subramaniam S, Science 348 1147-1151 (2015).
- PMID: 25953817
- EMDB entry EMD-2984 (image below, left)
- PDB entry 5a1a (image below, right)

At present, the PDB structure has been cited in 166 publications (56 reviews and 110 articles), the EMDB map in 48 publications and the EMPIAR dataset in 8 publications.
The dataset is 12.4 TB in size (until 2019 this was the biggest dataset in EMPIAR) and would not have been trivial to host or to download through ftp. Even with fast Aspera or Globus downloading from EMBL-EBI, this may take anywhere from a couple of hours to as much as a week (depending on the speed of the connection); with ftp it would have taken much longer.
Re-processing
By making the raw data publicly available in EMPIAR, other cryo-EM people have been able to download, analyse and re-process the data. At the time of publication, there was some uncertainty in the community about the claimed resolution. However, since the data was available, anyone with an interest could download and re-process the data and this has been done by quite a few groups. Some of these have deposited the resulting map in EMDB themselves (and sometimes also the model in the PDB). Two notable examples of this are:
- the original authors reprocessed their own data in 2018 and the resulting 1.9Å map is in EMDB as entry EMD-7770 (image below, left) and the model in the PDB as entry 6cvm (image below, right). This is described in paper 29754826.
-
Sjors Scheres is the author of the Relion software which is the most-used programme
for image processing in single-particle cryo-EM studies (~90% of all EMDB entries
released in 2019 used Relion). He has used the EMPIAR dataset on at least two occasions:
- To obtain a 2.2Å map using Relion version 2.0 in 2016 – this map is in EMDB as entry EMD-4116 (paper 27845625 ) (image below, left).
- To test version 3.0 of Relion, in 2018 the data was reprocessed to 1.9Å and the map deposited in EMDB as entry EMD-0153 (paper 30412051 ) (image below, right).


Other re-uses
Other examples that demonstrate how this EMPIAR entry has helped scientists develop new or improved (software) methods include:
-
Two cases of developing deep-learning methods:
- Case 1: Automated particle-picking using convolutional neuronal networks ( https://doi.org/10.1101/616169 )
- Case 2: Convolutional neural network for reducing false positives from particle picking ( https://doi.org/10.1107/S2052252518014392 )
-
Three examples of improving general cryo-EM processing software:
- Case 1: Development of RELION 2 by the Scheres group mentioned above ( https://doi.org/10.7554/eLife.18722 ).
- Case 2: Development of RELION 3 by the Scheres group mentioned above ( https://doi.org/10.7554/eLife.42166 ).
- Case 3: Development of cisTEM by Timothy Grant (while in the Grigorieff group at Janelia; https://doi.org/10.7554/eLife.35383 ). This is also a general image-processing software package for cryo-EM.
- One example of developing cloud-based processing: Set-up and use of RELION on Amazon Web Services ( https://doi.org/10.1016/j.jsb.2018.05.014 )
Additional types of re-use that have occurred for EMPIAR datasets (but not necessarily this one) include:
- Use in community challenges where data from EMPIAR is downloaded by multiple independent labs and re-processed using local procedures and software, and the resulting maps (and sometimes models built into those maps) are compared in a community meeting.
- Use in software tutorials and other training materials.
- Local use to train new students etc. in the processing of cryo-EM data.
- Validation of the published map or model obtained with a dataset.
Download statistics
Although we don’t track individual downloads, we have been able to determine that between 1 April 2018 and 1 December 2019 (20 months), 108,000 files from this entry have been downloaded by ftp and 127,000 through Aspera (Globus download statistics are not available). The entry itself contains 3079 files but users can choose to download any or all of these files. Assuming all users downloaded the entire dataset, this means that it has been downloaded ~75 times in this period. The entry was released in May 2016 and it seems not unlikely that the entry would have been downloaded at least as many times in the period May 2016-March 2018.
The statistics do not include files that have been downloaded through the EMPIAR mirror in Osaka.

