
|

|

|
|
 |
Contents |
 |
|
|

|

|

|

|

|

|

|



 229 a.a.
229 a.a.
|
 |

|

|

|

|

|

|

|

 1119 a.a.
1119 a.a.
|
 |

|

|

|

|

|

|

|

 1381 a.a.
1381 a.a.
|
 |

|

|

|

|

|

|

|

 95 a.a.
95 a.a.
|
 |

|

|

|

|

|

|

|

 345 a.a.
345 a.a.
|
 |

|

|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Transferase
|
 |
|
Title:
|
 |
Crystal structure of the t. Thermophilus RNA polymerase holoenzyme in complex with antibiotic sterptolydigin
|
|
Structure:
|
 |
DNA-directed RNA polymerase alpha chain. Chain: a, b, k, l. Synonym: rnap alpha subunit, transcriptase alpha chain, RNA polymerase alpha subunit. DNA-directed RNA polymerase beta chain. Chain: c, m. Synonym: rnap beta subunit, transcriptase beta chain, RNA polymerase beta subunit. DNA-directed RNA polymerase beta' chain.
|
|
Source:
|
 |
Thermus thermophilus. Organism_taxid: 274. Organism_taxid: 274
|
|
Biol. unit:
|
 |
 Hexamer (from
)
Hexamer (from
)
|
|
Resolution:
|
 |
|
2.40Å
|
R-factor:
|
0.230
|
R-free:
|
0.268
|
|
|
Authors:
|
 |
D.Temiakov,N.Zenkin,M.N.Vassylyeva,A.Perederina,T.H.Tahirov, M.Savkina,S.Zorov,V.Nikiforov,N.Igarashi,N.Matsugaki,S.Wakatsuki, K.Severinov,D.G.Vassylyev,Riken Structural Genomics/proteomics Initiative (Rsgi)
|
Key ref:

|
 |
D.Temiakov
et al.
(2005).
Structural basis of transcription inhibition by antibiotic streptolydigin.
Mol Cell,
19,
655-666.
PubMed id:
DOI:

|
 |
|
Date:
|
 |
|
02-Jul-05
|
Release date:
|
20-Sep-05
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
Q5SHR6
(RPOA_THET8) -
DNA-directed RNA polymerase subunit alpha from Thermus thermophilus (strain ATCC 27634 / DSM 579 / HB8)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
315 a.a.
229 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
Q8RQE9
(RPOB_THET8) -
DNA-directed RNA polymerase subunit beta from Thermus thermophilus (strain ATCC 27634 / DSM 579 / HB8)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
1119 a.a.
1119 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
Q8RQE8
(RPOC_THET8) -
DNA-directed RNA polymerase subunit beta' from Thermus thermophilus (strain ATCC 27634 / DSM 579 / HB8)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
1524 a.a.
1381 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|
 |
|
|
 |
 |
 |
 |
Enzyme class:
|
 |
Chains A, B, C, D, E, K, L, M, N, O:
E.C.2.7.7.6
- DNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
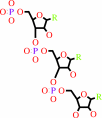 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Mol Cell
19:655-666
(2005)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structural basis of transcription inhibition by antibiotic streptolydigin.
|
|
D.Temiakov,
N.Zenkin,
M.N.Vassylyeva,
A.Perederina,
T.H.Tahirov,
E.Kashkina,
M.Savkina,
S.Zorov,
V.Nikiforov,
N.Igarashi,
N.Matsugaki,
S.Wakatsuki,
K.Severinov,
D.G.Vassylyev.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Streptolydigin (Stl) is a potent inhibitor of bacterial RNA polymerases (RNAPs).
The 2.4 A resolution structure of the Thermus thermophilus RNAP-Stl complex
showed that, in full agreement with the available genetic data, the inhibitor
binding site is located 20 A away from the RNAP active site and encompasses the
bridge helix and the trigger loop, two elements that are considered to be
crucial for RNAP catalytic center function. Structure-based biochemical
experiments revealed additional determinants of Stl binding and demonstrated
that Stl does not affect NTP substrate binding, DNA translocation, and
phosphodiester bond formation. The RNAP-Stl complex structure, its comparison
with the closely related substrate bound eukaryotic transcription elongation
complexes, and biochemical analysis suggest an inhibitory mechanism in which Stl
stabilizes catalytically inactive (preinsertion) substrate bound transcription
intermediate, thereby blocking structural isomerization of RNAP to an active
configuration. The results provide a basis for a design of new antibiotics
utilizing the Stl-like mechanism.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 1.
Figure 1. The RNAP-Stl Complex Structure
|
 |
Figure 5.
Figure 5. Substrate Loading in the IS and Plausible
Mechanism of Stl Action
|
 |
|
|

|
| |
The above figures are
reprinted
by permission from Cell Press:
Mol Cell
(2005,
19,
655-666)
copyright 2005.
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
S.R.Kennedy,
and
D.A.Erie
(2011).
Templated nucleoside triphosphate binding to a noncatalytic site on RNA polymerase regulates transcription.
|
| |
Proc Natl Acad Sci U S A,
108,
6079-6084.
|
 |
|
|
|
|
 |
C.D.Kaplan
(2010).
The architecture of RNA polymerase fidelity.
|
| |
BMC Biol,
8,
85.
|
 |
|
|
|
|
 |
J.C.Carlson,
J.L.Fortman,
Y.Anzai,
S.Li,
D.A.Burr,
and
D.H.Sherman
(2010).
Identification of the tirandamycin biosynthetic gene cluster from Streptomyces sp. 307-9.
|
| |
Chembiochem,
11,
564-572.
|
 |
|
|
|
|
 |
J.Zhang,
M.Palangat,
and
R.Landick
(2010).
Role of the RNA polymerase trigger loop in catalysis and pausing.
|
| |
Nat Struct Mol Biol,
17,
99.
|
 |
|
|
|
|
 |
M.Sánchez-Hidalgo,
L.E.Núñez,
C.Méndez,
and
J.A.Salas
(2010).
Involvement of the beta subunit of RNA polymerase in resistance to streptolydigin and streptovaricin in the producer organisms Streptomyces lydicus and Streptomyces spectabilis.
|
| |
Antimicrob Agents Chemother,
54,
1684-1692.
|
 |
|
|
|
|
 |
S.Tagami,
S.Sekine,
T.Kumarevel,
N.Hino,
Y.Murayama,
S.Kamegamori,
M.Yamamoto,
K.Sakamoto,
and
S.Yokoyama
(2010).
Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein.
|
| |
Nature,
468,
978-982.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
W.J.Lane,
and
S.A.Darst
(2010).
Molecular evolution of multisubunit RNA polymerases: structural analysis.
|
| |
J Mol Biol,
395,
686-704.
|
 |
|
|
|
|
 |
Y.Yuzenkova,
A.Bochkareva,
V.R.Tadigotla,
M.Roghanian,
S.Zorov,
K.Severinov,
and
N.Zenkin
(2010).
Stepwise mechanism for transcription fidelity.
|
| |
BMC Biol,
8,
54.
|
 |
|
|
|
|
 |
Y.Yuzenkova,
and
N.Zenkin
(2010).
Central role of the RNA polymerase trigger loop in intrinsic RNA hydrolysis.
|
| |
Proc Natl Acad Sci U S A,
107,
10878-10883.
|
 |
|
|
|
|
 |
C.Olano,
C.Gómez,
M.Pérez,
M.Palomino,
A.Pineda-Lucena,
R.J.Carbajo,
A.F.Braña,
C.Méndez,
and
J.A.Salas
(2009).
Deciphering biosynthesis of the RNA polymerase inhibitor streptolydigin and generation of glycosylated derivatives.
|
| |
Chem Biol,
16,
1031-1044.
|
 |
|
|
|
|
 |
D.G.Vassylyev
(2009).
Elongation by RNA polymerase: a race through roadblocks.
|
| |
Curr Opin Struct Biol,
19,
691-700.
|
 |
|
|
|
|
 |
E.Nudler
(2009).
RNA polymerase active center: the molecular engine of transcription.
|
| |
Annu Rev Biochem,
78,
335-361.
|
 |
|
|
|
|
 |
G.A.Belogurov,
M.N.Vassylyeva,
A.Sevostyanova,
J.R.Appleman,
A.X.Xiang,
R.Lira,
S.E.Webber,
S.Klyuyev,
E.Nudler,
I.Artsimovitch,
and
D.G.Vassylyev
(2009).
Transcription inactivation through local refolding of the RNA polymerase structure.
|
| |
Nature,
457,
332-335.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
J.C.Carlson,
S.Li,
D.A.Burr,
and
D.H.Sherman
(2009).
Isolation and characterization of tirandamycins from a marine-derived Streptomyces sp.
|
| |
J Nat Prod,
72,
2076-2079.
|
 |
|
|
|
|
 |
M.X.Ho,
B.P.Hudson,
K.Das,
E.Arnold,
and
R.H.Ebright
(2009).
Structures of RNA polymerase-antibiotic complexes.
|
| |
Curr Opin Struct Biol,
19,
715-723.
|
 |
|
|
|
|
 |
N.Miropolskaya,
I.Artsimovitch,
S.Klimasauskas,
V.Nikiforov,
and
A.Kulbachinskiy
(2009).
Allosteric control of catalysis by the F loop of RNA polymerase.
|
| |
Proc Natl Acad Sci U S A,
106,
18942-18947.
|
 |
|
|
|
|
 |
C.D.Kaplan,
K.M.Larsson,
and
R.D.Kornberg
(2008).
The RNA polymerase II trigger loop functions in substrate selection and is directly targeted by alpha-amanitin.
|
| |
Mol Cell,
30,
547-556.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
D.Dutta,
J.Chalissery,
and
R.Sen
(2008).
Transcription termination factor rho prefers catalytically active elongation complexes for releasing RNA.
|
| |
J Biol Chem,
283,
20243-20251.
|
 |
|
|
|
|
 |
L.Tan,
S.Wiesler,
D.Trzaska,
H.C.Carney,
and
R.O.Weinzierl
(2008).
Bridge helix and trigger loop perturbations generate superactive RNA polymerases.
|
| |
J Biol,
7,
40.
|
 |
|
|
|
|
 |
S.Borukhov,
and
E.Nudler
(2008).
RNA polymerase: the vehicle of transcription.
|
| |
Trends Microbiol,
16,
126-134.
|
 |
|
|
|
|
 |
D.G.Vassylyev,
M.N.Vassylyeva,
J.Zhang,
M.Palangat,
I.Artsimovitch,
and
R.Landick
(2007).
Structural basis for substrate loading in bacterial RNA polymerase.
|
| |
Nature,
448,
163-168.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
I.Toulokhonov,
J.Zhang,
M.Palangat,
and
R.Landick
(2007).
A central role of the RNA polymerase trigger loop in active-site rearrangement during transcriptional pausing.
|
| |
Mol Cell,
27,
406-419.
|
 |
|
|
|
|
 |
M.N.Vassylyeva,
V.Svetlov,
A.D.Dearborn,
S.Klyuyev,
I.Artsimovitch,
and
D.G.Vassylyev
(2007).
The carboxy-terminal coiled-coil of the RNA polymerase beta'-subunit is the main binding site for Gre factors.
|
| |
EMBO Rep,
8,
1038-1043.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.Kyzer,
K.S.Ha,
R.Landick,
and
M.Palangat
(2007).
Direct versus limited-step reconstitution reveals key features of an RNA hairpin-stabilized paused transcription complex.
|
| |
J Biol Chem,
282,
19020-19028.
|
 |
|
|
|
|
 |
V.Svetlov,
G.A.Belogurov,
E.Shabrova,
D.G.Vassylyev,
and
I.Artsimovitch
(2007).
Allosteric control of the RNA polymerase by the elongation factor RfaH.
|
| |
Nucleic Acids Res,
35,
5694-5705.
|
 |
|
|
|
|
 |
Y.Xiong,
and
Z.F.Burton
(2007).
A tunable ratchet driving human RNA polymerase II translocation adjusted by accurately templated nucleoside triphosphates loaded at downstream sites and by elongation factors.
|
| |
J Biol Chem,
282,
36582-36592.
|
 |
|
|
|
|
 |
D.Wang,
D.A.Bushnell,
K.D.Westover,
C.D.Kaplan,
and
R.D.Kornberg
(2006).
Structural basis of transcription: role of the trigger loop in substrate specificity and catalysis.
|
| |
Cell,
127,
941-954.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
E.Kashkina,
M.Anikin,
T.H.Tahirov,
S.N.Kochetkov,
D.G.Vassylyev,
and
D.Temiakov
(2006).
Elongation complexes of Thermus thermophilus RNA polymerase that possess distinct translocation conformations.
|
| |
Nucleic Acids Res,
34,
4036-4045.
|
 |
|
|
|
|
 |
N.Zenkin,
Y.Yuzenkova,
and
K.Severinov
(2006).
Transcript-assisted transcriptional proofreading.
|
| |
Science,
313,
518-520.
|
 |
|
|
|
|
 |
S.F.Holmes,
T.J.Santangelo,
C.K.Cunningham,
J.W.Roberts,
and
D.A.Erie
(2006).
Kinetic investigation of Escherichia coli RNA polymerase mutants that influence nucleotide discrimination and transcription fidelity.
|
| |
J Biol Chem,
281,
18677-18683.
|
 |
|
|
|
|
 |
V.Trinh,
M.F.Langelier,
J.Archambault,
and
B.Coulombe
(2006).
Structural perspective on mutations affecting the function of multisubunit RNA polymerases.
|
| |
Microbiol Mol Biol Rev,
70,
12-36.
|
 |
|
|
|
|
 |
D.G.Vassylyev,
and
I.Artsimovitch
(2005).
Tracking RNA polymerase, one step at a time.
|
| |
Cell,
123,
977-979.
|
 |
|
|
|
|
 |
D.G.Vassylyev,
V.Svetlov,
M.N.Vassylyeva,
A.Perederina,
N.Igarashi,
N.Matsugaki,
S.Wakatsuki,
and
I.Artsimovitch
(2005).
Structural basis for transcription inhibition by tagetitoxin.
|
| |
Nat Struct Mol Biol,
12,
1086-1093.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
E.A.Abbondanzieri,
W.J.Greenleaf,
J.W.Shaevitz,
R.Landick,
and
S.M.Block
(2005).
Direct observation of base-pair stepping by RNA polymerase.
|
| |
Nature,
438,
460-465.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|

| |

