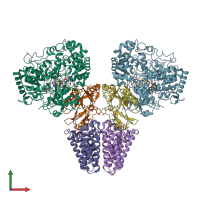
Assemblies
Multimeric state:
hetero hexamer
Accessible surface area:
76396.66 Å2
Buried surface area:
27348.25 Å2
Dissociation area:
1,507.59
Å2
Dissociation energy (ΔGdiss):
7.91
kcal/mol
Dissociation entropy (TΔSdiss):
16.76
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-180887
Macromolecules
Chains: A, E
Length: 765 amino acids
Theoretical weight: 86.74 KDa
Source organism: Thermus thermophilus HB27
UniProt:
Pfam:
Length: 765 amino acids
Theoretical weight: 86.74 KDa
Source organism: Thermus thermophilus HB27
UniProt:
- Canonical:
 Q72LA4 (Residues: 1-765; Coverage: 100%)
Q72LA4 (Residues: 1-765; Coverage: 100%)
Pfam:
- Molybdopterin oxidoreductase Fe4S4 domain
- Molybdopterin oxidoreductase
- Molydopterin dinucleotide binding domain
- Twin-arginine translocation pathway, signal sequence
- Twin-arginine translocation pathway, signal sequence, bacterial/archaeal
- Molybdopterin oxidoreductase, 4Fe-4S domain
- Molybdopterin oxidoreductase
- Aspartate decarboxylase-like domain superfamily
- Molybdopterin dinucleotide-binding domain
Chains: B, F
Length: 195 amino acids
Theoretical weight: 21.41 KDa
Source organism: Thermus thermophilus HB27
UniProt:
Pfam:
InterPro:
Length: 195 amino acids
Theoretical weight: 21.41 KDa
Source organism: Thermus thermophilus HB27
UniProt:
- Canonical:
 Q72LA5 (Residues: 1-195; Coverage: 100%)
Q72LA5 (Residues: 1-195; Coverage: 100%)
Pfam:
InterPro:
- 4Fe-4S ferredoxin-type, iron-sulphur binding domain
- 7Fe ferredoxin
- 4Fe-4S ferredoxin, iron-sulphur binding, conserved site
Chains: C, G
Length: 253 amino acids
Theoretical weight: 27.62 KDa
Source organism: Thermus thermophilus HB27
UniProt:
Pfam: Polysulfide reductase
InterPro: Polysulfide reductase
CATH: Formate dehydrogenase/DMSO reductase domain
Length: 253 amino acids
Theoretical weight: 27.62 KDa
Source organism: Thermus thermophilus HB27
UniProt:
- Canonical:
 Q72LA6 (Residues: 1-253; Coverage: 100%)
Q72LA6 (Residues: 1-253; Coverage: 100%)
Pfam: Polysulfide reductase
InterPro: Polysulfide reductase
CATH: Formate dehydrogenase/DMSO reductase domain