Assemblies
Assembly Name:
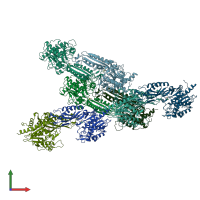
Peptidase M20 dimerisation domain-containing protein
Multimeric state:
homo dimer
Accessible surface area:
33162.05 Å2
Buried surface area:
5739.11 Å2
Dissociation area:
2,459.52
Å2
Dissociation energy (ΔGdiss):
17.8
kcal/mol
Dissociation entropy (TΔSdiss):
14.68
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188776
Assembly Name:
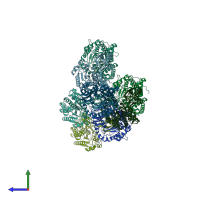
Peptidase M20 dimerisation domain-containing protein
Multimeric state:
homo dimer
Accessible surface area:
33371.72 Å2
Buried surface area:
5811.05 Å2
Dissociation area:
2,482.47
Å2
Dissociation energy (ΔGdiss):
20.51
kcal/mol
Dissociation entropy (TΔSdiss):
14.7
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188776
Assembly Name:
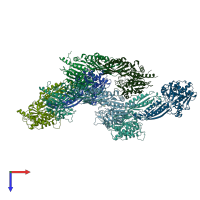
Peptidase M20 dimerisation domain-containing protein
Multimeric state:
homo dimer
Accessible surface area:
33619.21 Å2
Buried surface area:
5817.59 Å2
Dissociation area:
2,495.7
Å2
Dissociation energy (ΔGdiss):
22.46
kcal/mol
Dissociation entropy (TΔSdiss):
14.69
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188776
Assembly Name:
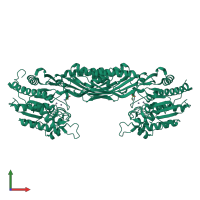
Peptidase M20 dimerisation domain-containing protein
Multimeric state:
homo dimer
Accessible surface area:
33419.65 Å2
Buried surface area:
5739.53 Å2
Dissociation area:
2,544.52
Å2
Dissociation energy (ΔGdiss):
18.56
kcal/mol
Dissociation entropy (TΔSdiss):
14.71
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188776
Macromolecules
Chains: A, B, C, D, E, F, G, H
Length: 462 amino acids
Theoretical weight: 50.73 KDa
Source organism: Lachancea kluyveri
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
SCOP:
Length: 462 amino acids
Theoretical weight: 50.73 KDa
Source organism: Lachancea kluyveri
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q96W94 (Residues: 2-455; Coverage: 100%)
Q96W94 (Residues: 2-455; Coverage: 100%)
Pfam:
InterPro:
- Amidase, carbamoylase-type
- Peptidase M20
- Bacterial exopeptidase dimerisation domain
- Peptidase M20, dimerisation domain
SCOP: