 |
PDBsum entry 4bbs

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transcription
|
PDB id
|
|

|

|
4bbs
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|

|

|

|

|

|
 1419 a.a.
1419 a.a.
|
 |

|

|

|

|

|

|

|
 1150 a.a.
1150 a.a.
|
 |

|

|

|

|

|

|

|
 266 a.a.
266 a.a.
|
 |

|

|

|

|

|

|

|
 178 a.a.
178 a.a.
|
 |

|

|

|

|

|

|

|
 214 a.a.
214 a.a.
|
 |

|

|

|

|

|

|

|
 87 a.a.
87 a.a.
|
 |

|

|

|

|

|

|

|
 171 a.a.
171 a.a.
|
 |

|

|

|

|

|

|

|
 134 a.a.
134 a.a.
|
 |

|

|

|

|

|

|

|
 119 a.a.
119 a.a.
|
 |

|

|

|

|

|

|

|
 65 a.a.
65 a.a.
|
 |

|

|

|

|

|

|

|
 114 a.a.
114 a.a.
|
 |

|

|

|

|

|

|

|
 44 a.a.
44 a.a.
|
 |

|

|

|

|

|

|

|
 189 a.a.
189 a.a.
|
 |

|

|
|
|
|
|
|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Transcription
|
 |
|
Title:
|
 |
Structure of an initially transcribing RNA polymerase ii-tfiib complex
|
|
Structure:
|
 |
DNA-directed RNA polymerase ii subunit rpb1. Chain: a. Synonym: RNA polymerase ii subunit 1, RNA polymerase ii subunit b1, DNA-directed RNA polymerase iii largest subunit, RNA polymerase ii subunit b220, rpb1. DNA-directed RNA polymerase ii subunit rpb2. Chain: b. Synonym: RNA polymerase ii subunit 2, b150, rpb2, DNA-directed RNA polymerase ii 140 kda polypeptide.
|
|
Source:
|
 |
Saccharomyces cerevisiae. Baker's yeast. Organism_taxid: 4932. Expressed in: escherichia coli. Expression_system_taxid: 562. Synthetic: yes. Organism_taxid: 4932
|
|
Resolution:
|
 |
|
3.60Å
|
R-factor:
|
0.186
|
R-free:
|
0.225
|
|
|
Authors:
|
 |
S.Sainsbury,J.Niesser,P.Cramer
|
|
Key ref:
|
 |
S.Sainsbury
et al.
(2013).
Structure and function of the initially transcribing RNA polymerase II-TFIIB complex.
Nature,
493,
437-440.
PubMed id:
|
 |
|
Date:
|
 |
|
27-Sep-12
|
Release date:
|
14-Nov-12
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P04050
(RPB1_YEAST) -
DNA-directed RNA polymerase II subunit RPB1 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
1733 a.a.
1419 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P08518
(RPB2_YEAST) -
DNA-directed RNA polymerase II subunit RPB2 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
1224 a.a.
1150 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P16370
(RPB3_YEAST) -
DNA-directed RNA polymerase II subunit RPB3 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
318 a.a.
266 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P20433
(RPB4_YEAST) -
DNA-directed RNA polymerase II subunit RPB4 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
221 a.a.
178 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P20434
(RPAB1_YEAST) -
DNA-directed RNA polymerases I, II, and III subunit RPABC1 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
215 a.a.
214 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P20435
(RPAB2_YEAST) -
DNA-directed RNA polymerases I, II, and III subunit RPABC2 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
155 a.a.
87 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P34087
(RPB7_YEAST) -
DNA-directed RNA polymerase II subunit RPB7 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
171 a.a.
171 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P20436
(RPAB3_YEAST) -
DNA-directed RNA polymerases I, II, and III subunit RPABC3 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
146 a.a.
134 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P27999
(RPB9_YEAST) -
DNA-directed RNA polymerase II subunit RPB9 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
122 a.a.
119 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P22139
(RPAB5_YEAST) -
DNA-directed RNA polymerases I, II, and III subunit RPABC5 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
70 a.a.
65 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|

|

|
|
P38902
(RPB11_YEAST) -
DNA-directed RNA polymerase II subunit RPB11 from Saccharomyces cerevisiae (strain ATCC 204508 / S288c)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
120 a.a.
114 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|
 |
|
|
 |
 |
 |
 |
Enzyme class:
|
 |
Chains A, B:
E.C.2.7.7.6
- DNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
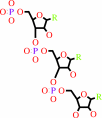 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
Nature
493:437-440
(2013)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structure and function of the initially transcribing RNA polymerase II-TFIIB complex.
|
|
S.Sainsbury,
J.Niesser,
P.Cramer.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The general transcription factor (TF) IIB is required for RNA polymerase (Pol)
II initiation and extends with its B-reader element into the Pol II active
centre cleft. Low-resolution structures of the Pol II-TFIIB complex indicated
how TFIIB functions in DNA recruitment, but they lacked nucleic acids and half
of the B-reader, leaving other TFIIB functions enigmatic. Here we report crystal
structures of the Pol II-TFIIB complex from the yeast Saccharomyces cerevisiae
at 3.4 Å resolution and of an initially transcribing complex that
additionally contains the DNA template and a 6-nucleotide RNA product. The
structures reveal the entire B-reader and protein-nucleic acid interactions, and
together with functional data lead to a more complete understanding of
transcription initiation. TFIIB partially closes the polymerase cleft to
position DNA and assist in its opening. The B-reader does not reach the active
site but binds the DNA template strand upstream to assist in the recognition of
the initiator sequence and in positioning the transcription start site. TFIIB
rearranges active-site residues, induces binding of the catalytic metal ion B,
and stimulates initial RNA synthesis allosterically. TFIIB then prevents the
emerging DNA-RNA hybrid duplex from tilting, which would impair RNA synthesis.
When the RNA grows beyond 6 nucleotides, it is separated from DNA and is
directed to its exit tunnel by the B-reader loop. Once the RNA grows to 12-13
nucleotides, it clashes with TFIIB, triggering TFIIB displacement and elongation
complex formation. Similar mechanisms may underlie all cellular transcription
because all eukaryotic and archaeal RNA polymerases use TFIIB-like factors, and
the bacterial initiation factor sigma has TFIIB-like topology and contains the
loop region 3.2 that resembles the B-reader loop in location, charge and
function. TFIIB and its counterparts may thus account for the two fundamental
properties that distinguish RNA from DNA polymerases: primer-independent chain
initiation and product separation from the template.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |

| | |

