 |
PDBsum entry 3kk3

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase/DNA
|
PDB id
|
|

|

|
3kk3
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
Chains A, B:
E.C.2.7.7.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
Chains A, B:
E.C.2.7.7.49
- RNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
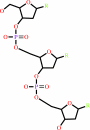 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
Chains A, B:
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
Chains A, B:
E.C.3.1.-.-
|
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
Chains A, B:
E.C.3.1.13.2
- exoribonuclease H.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Exonucleolytic cleavage to 5'-phosphomonoester oligonucleotides in both 5'- to 3'- and 3'- to 5'-directions.
|
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
Chains A, B:
E.C.3.1.26.13
- retroviral ribonuclease H.
|
|
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
Chains A, B:
E.C.3.4.23.16
- HIV-1 retropepsin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Specific for a P1 residue that is hydrophobic, and P1' variable, but often Pro.
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
J Mol Biol
397:967-978
(2010)
|
|
PubMed id:
|
|
|
|

|
| |
|
Visualizing the molecular interactions of a nucleotide analog, GS-9148, with HIV-1 reverse transcriptase-DNA complex.
|
|
E.B.Lansdon,
D.Samuel,
L.Lagpacan,
K.M.Brendza,
K.L.White,
M.Hung,
X.Liu,
C.G.Boojamra,
R.L.Mackman,
T.Cihlar,
A.S.Ray,
M.E.McGrath,
S.Swaminathan.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
GS-9148
([5-(6-amino-purin-9-yl)-4-fluoro-2,5-dihydro-furan-2-yloxymethyl]-phosphonic
acid) is a dAMP (2'-deoxyadenosine monophosphate) analog that maintains its
antiviral activity against drug-resistant HIV. Crystal structures for HIV-1
reverse transcriptase (RT) bound to double-stranded DNA, ternary complexes with
either GS-9148-diphosphate or 2'-deoxyadenosine triphosphate (dATP), and a
post-incorporation structure with GS-9148 translocated to the priming site were
obtained to gain insight into the mechanism of RT inhibition. The binding of
either GS-9148-diphosphate or dATP to the binary RT-DNA complex resulted in the
fingers subdomain closing around the incoming substrate. This produced up to a 9
A shift in the tips of the fingers subdomain as it closed toward the palm and
thumb subdomains. GS-9148-diphosphate shows a similar binding mode as dATP in
the nucleotide-binding site. Residues whose mutations confer resistance to
nucleotide/nucleoside RT inhibitors, such as M184, Y115, L74, and K65, show
little to no shift in orientation whether GS-9148-diphosphate or dATP is bound.
One difference observed in binding is the position of the central ring. The
dihydrofuran ring of GS-9148-diphosphate interacts with the aromatic side chain
of Y115 more than does the ribose ring of dATP, possibly picking up a favorable
pi-pi interaction. The ability of GS-9148-diphosphate to mimic the active-site
contacts of dATP may explain its effective inhibition of RT and maintained
activity against resistance mutations. Interestingly, the 2'-fluoro moiety of
GS-9148-diphosphate was found in close proximity to the Q151 side chain,
potentially explaining the observed moderately reduced susceptibly to GS-9148
conferred by Q151M mutation.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
M.Lapkouski,
L.Tian,
J.T.Miller,
S.F.Le Grice,
and
W.Yang
(2013).
Complexes of HIV-1 RT, NNRTI and RNA/DNA hybrid reveal a structure compatible with RNA degradation.
|
| |
Nat Struct Mol Biol,
20,
230-236.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Das,
S.E.Martinez,
J.D.Bauman,
and
E.Arnold
(2012).
HIV-1 reverse transcriptase complex with DNA and nevirapine reveals non-nucleoside inhibition mechanism.
|
| |
Nat Struct Mol Biol,
19,
253-259.
|
 |
|
PDB codes:
|
 |
|
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |
|

