Assemblies
Assembly Name:
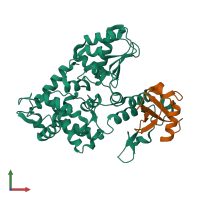
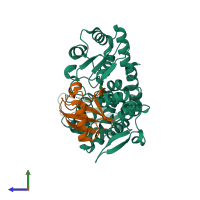
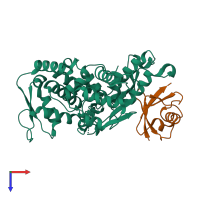
E3 ubiquitin-protein ligase Itchy homolog and Ubiquitin
Multimeric state:
hetero dimer
Accessible surface area:
21981.42 Å2
Buried surface area:
1462.93 Å2
Dissociation area:
731.46
Å2
Dissociation energy (ΔGdiss):
-0.6
kcal/mol
Dissociation entropy (TΔSdiss):
10.77
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-143398
Assembly Name:
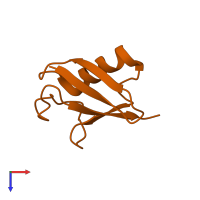
Ubiquitin
Multimeric state:
monomeric
Accessible surface area:
4719.97 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-143327
Macromolecules
Chain: A
Length: 394 amino acids
Theoretical weight: 47.03 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: HECT-domain (ubiquitin-transferase)
InterPro:
Length: 394 amino acids
Theoretical weight: 47.03 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q96J02 (Residues: 524-899; Coverage: 42%)
Q96J02 (Residues: 524-899; Coverage: 42%)
Pfam: HECT-domain (ubiquitin-transferase)
InterPro:
Chains: B, C
Length: 84 amino acids
Theoretical weight: 9.77 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ubiquitin family
InterPro:
Length: 84 amino acids
Theoretical weight: 9.77 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P0CG48 (Residues: 533-607; Coverage: 11%)
P0CG48 (Residues: 533-607; Coverage: 11%)
Pfam: Ubiquitin family
InterPro: