Assemblies
Assembly Name:
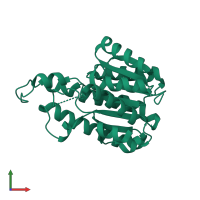
Dehydrogenase/reductase SDR family member 1
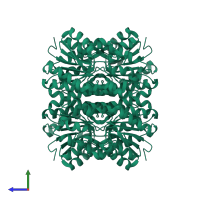
Multimeric state:
homo dimer
Accessible surface area:
18348.8 Å2
Buried surface area:
3505.84 Å2
Dissociation area:
1,752.92
Å2
Dissociation energy (ΔGdiss):
23.29
kcal/mol
Dissociation entropy (TΔSdiss):
13.01
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188619
Assembly Name:
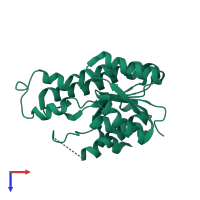
Dehydrogenase/reductase SDR family member 1
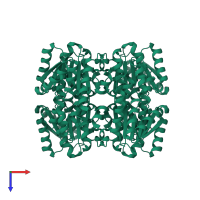
Multimeric state:
homo dimer
Accessible surface area:
20117.74 Å2
Buried surface area:
1736.89 Å2
Dissociation area:
868.45
Å2
Dissociation energy (ΔGdiss):
2.65
kcal/mol
Dissociation entropy (TΔSdiss):
12.76
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-188619
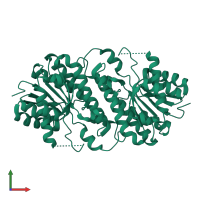
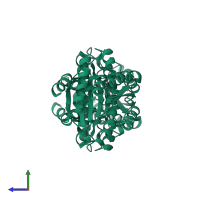
Assembly Name:
Dehydrogenase/reductase SDR family member 1
Multimeric state:
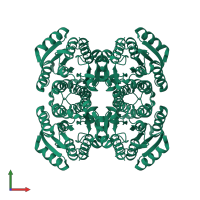
homo tetramer
Accessible surface area:
33169.05 Å2
Buried surface area:
10540.22 Å2
Dissociation area:
1,764.27
Å2
Dissociation energy (ΔGdiss):
16.12
kcal/mol
Dissociation entropy (TΔSdiss):
14.31
kcal/mol
Symmetry number:
4
PDBe Complex ID:
PDB-CPX-188620
Macromolecules
Chain: A
Length: 260 amino acids
Theoretical weight: 28.04 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: short chain dehydrogenase
InterPro:
CATH: NAD(P)-binding Rossmann-like Domain
Length: 260 amino acids
Theoretical weight: 28.04 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q96LJ7 (Residues: 3-262; Coverage: 83%)
Q96LJ7 (Residues: 3-262; Coverage: 83%)
Pfam: short chain dehydrogenase
InterPro:
CATH: NAD(P)-binding Rossmann-like Domain