Assemblies
Assembly Name:
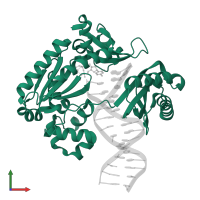
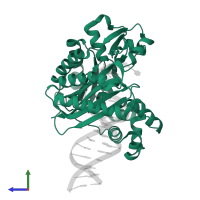
DNA polymerase IV and DNA
Multimeric state:
hetero trimer
Accessible surface area:
19695.17 Å2
Buried surface area:
5186.55 Å2
Dissociation area:
1,172.12
Å2
Dissociation energy (ΔGdiss):
33.88
kcal/mol
Dissociation entropy (TΔSdiss):
10.59
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-238519
Assembly Name:
DNA polymerase IV and DNA
Multimeric state:
hetero trimer
Accessible surface area:
19866.71 Å2
Buried surface area:
5127.28 Å2
Dissociation area:
1,168.36
Å2
Dissociation energy (ΔGdiss):
32.71
kcal/mol
Dissociation entropy (TΔSdiss):
10.59
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-238519
Macromolecules
Chains: A, B
Length: 361 amino acids
Theoretical weight: 41.12 KDa
Source organisms: Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 361 amino acids
Theoretical weight: 41.12 KDa
Source organisms: Expression system: Escherichia coli
UniProt:
- Canonical:
 Q4JB80 (Residues: 1-231; Coverage: 65%)
Q4JB80 (Residues: 1-231; Coverage: 65%) - Canonical:
 Q97W02 (Residues: 231-352; Coverage: 35%)
nullnull
Q97W02 (Residues: 231-352; Coverage: 35%)
nullnull
Pfam:
InterPro:
- DNA polymerase IV
- DNA/RNA polymerase superfamily
- UmuC domain
- Reverse transcriptase/Diguanylate cyclase domain
- DNA polymerase type-Y, HhH motif
- DNA polymerase, Y-family, little finger domain superfamily
- DNA polymerase, Y-family, little finger domain
Name:
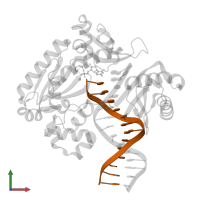
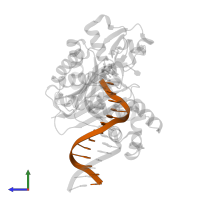
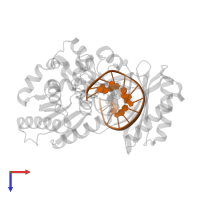
DNA (5'-D(*GP*GP*CP*AP*CP*TP*GP*AP*TP*CP*GP*GP*C)-3')
Representative chains: E, P
Length: 13 nucleotides
Theoretical weight: 3.99 KDa
Representative chains: E, P
Length: 13 nucleotides
Theoretical weight: 3.99 KDa
Name:
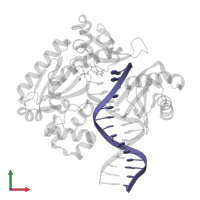
DNA (5'-D(*TP*TP*AP*CP*GP*GP*CP*CP*GP*AP*TP*CP*AP*GP*TP*GP*CP*C)-3')
Representative chains: F, T
Length: 18 nucleotides
Theoretical weight: 5.49 KDa
Representative chains: F, T
Length: 18 nucleotides
Theoretical weight: 5.49 KDa