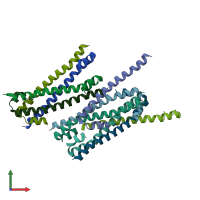Assemblies
Assembly Name:
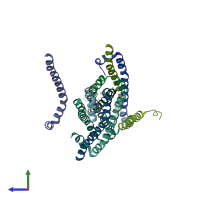
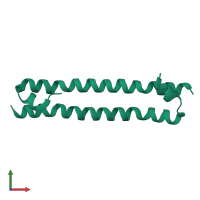
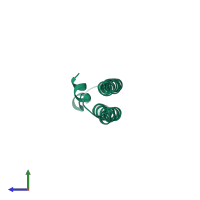
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7414.01 Å2
Buried surface area:
2861.43 Å2
Dissociation area:
1,430.72
Å2
Dissociation energy (ΔGdiss):
24.23
kcal/mol
Dissociation entropy (TΔSdiss):
10.9
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Assembly Name:
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7358.84 Å2
Buried surface area:
2933.59 Å2
Dissociation area:
1,466.8
Å2
Dissociation energy (ΔGdiss):
23.75
kcal/mol
Dissociation entropy (TΔSdiss):
10.89
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Assembly Name:
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7554.75 Å2
Buried surface area:
2799.26 Å2
Dissociation area:
1,399.63
Å2
Dissociation energy (ΔGdiss):
24.14
kcal/mol
Dissociation entropy (TΔSdiss):
10.85
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Assembly Name:
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7647.53 Å2
Buried surface area:
2849.89 Å2
Dissociation area:
1,424.95
Å2
Dissociation energy (ΔGdiss):
21.41
kcal/mol
Dissociation entropy (TΔSdiss):
10.86
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Assembly Name:
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7911.27 Å2
Buried surface area:
2597.92 Å2
Dissociation area:
1,298.96
Å2
Dissociation energy (ΔGdiss):
19.27
kcal/mol
Dissociation entropy (TΔSdiss):
10.89
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Assembly Name:
Serine/threonine-protein kinase 3 20kDa subunit
Multimeric state:
homo dimer
Accessible surface area:
7358.15 Å2
Buried surface area:
2484.13 Å2
Dissociation area:
1,242.06
Å2
Dissociation energy (ΔGdiss):
19.85
kcal/mol
Dissociation entropy (TΔSdiss):
10.55
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-171593
Macromolecules
Chains: A, B, C, D, E, F, G, H, I, J
Length: 51 amino acids
Theoretical weight: 6.17 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: C terminal SARAH domain of Mst1
InterPro:
CATH: p53-like tetramerisation domain
Length: 51 amino acids
Theoretical weight: 6.17 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q13188 (Residues: 436-484; Coverage: 10%)
Q13188 (Residues: 436-484; Coverage: 10%)
Pfam: C terminal SARAH domain of Mst1
InterPro:
CATH: p53-like tetramerisation domain