




 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots





 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots
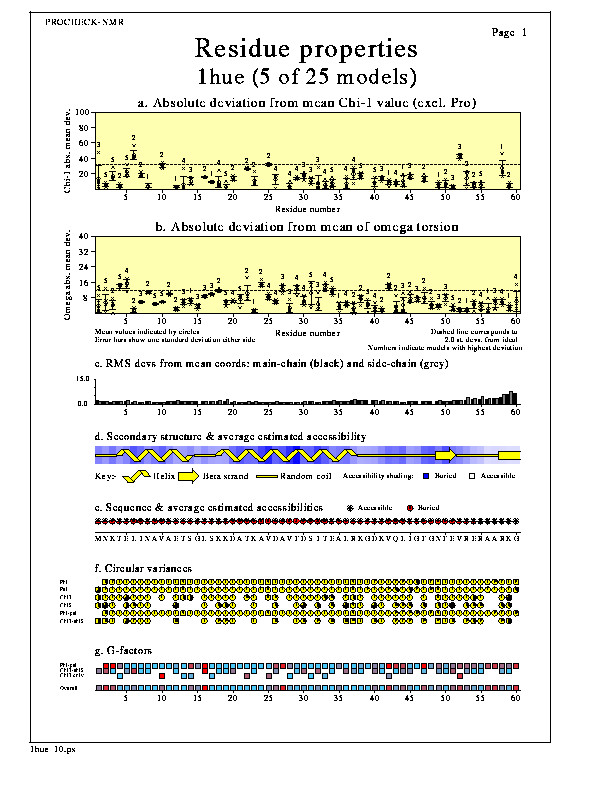
The Ensemble geometry plot shows how the protein's geometrical properties vary along its sequence. This gives a visualization of which regions appear to have consistently poor geometry (perhaps because they are poorly defined) and which have more normal geometry.
The properties plotted are:-
The first two graphs at the top of the page, can be selected from 3 possibles by the user. The two default graphs, which are plotted when you first run PROCHECK-NMR, are the first two of:-
For each graph, the values for each residue in each model are plotted as individual crosses. The mean and standard deviation values for a given residue are indicated by the circle and bars, respectively. The model whose value is the highest is indicated by the model-number printed above the highest mark.
The dashed line corresponds to 2.0 standard deviations from the ideal, so an excess of points above the line suggest possible problems with the geometry.
The RMS deviations plot shows a histogram of the rms deviations from the mean coordinates of the main-chain atoms (black bars) and the sidechain atoms (grey bars). The mean coordinates are currently calculated simply by averaging each atom's coordinates across the whole ensemble.
The secondary structure plot shows a schematic representation of the Kabsch & Sander (1983) secondary structure assignments. The key just below the picture shows which structure is which. Beta strands are taken to include all residues with a Kabsch & Sander assignment of E, helices corresponds to both H and G assignments, while everything else is taken to be random coil.
The shading behind the schematic picture gives an approximation to the residue accessibilities. The approximation is a fairly crude one, being based on each residue's Ooi number (Nishikawa & Ooi, 1986). An Ooi number is a count of the number of other Calpha atoms within a radius of, in this case, 14Å of the given residue's own Calpha. Although crude, this does give a good impression of which parts of the structure are buried and which are exposed on the surface. Future versions of PROCHECK will include an accurate calculation of residue accessibility.
The next section shows the sequence of the structure (using the 20 standard amino-acid codes) and a set of markers that give a schematic representation of the estimated accessibility. The latter is calculated as above, but here is show such that the darker the symbol the higher the accessibility.
The dials give a schematic representation of each residue's circular variance values for its phi, psi, chi-1 and chi-2 angles, and for its phi-psi and chi1-chi2 combinations. The larger the black segment on the dial, the higher the circular variance, and hence the wider the spread of the corresponding dihedral angle distribution.
Regions with many black, or near-black, dials correspond to regions where there is a large variability in the residue's tordion angles across the models in the ensemble. These may correspond to highly mobile or poorly defined regions such as loops, or may need further investigation.
The shaded squares give a schematic representation of each residue's G-factor values for its phi-psi and chi1-chi2 and chi-1 dihedral angles. (Note that the chi-1 G-factors are shown only for those residues that do not have a chi-2, and hence no chi1-chi2 G-factor).
Regions with many dark squares correspond to regions where the dihedral angles are "unusual", as defined by a low (or negative) G-factor. These may correspond to highly mobile or poorly defined regions such as loops, or may need further investigation.
The main options for the Ensemble geometry are:-
These options can be altered by editing the parameter file, procheck_nmr.prm, as described here.





 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots