The circular variance provides a measure of the spread of a
set of dihedral angles. It is applied here to each residue's
distribution of
![]() ,
,
![]() ,
,
![]() and
and
![]() angles across all the members of the NMR ensemble.
angles across all the members of the NMR ensemble.
So, for example, it can provide a measure of how tightly or loosely a given
residue's
![]() torsion angles cluster together across the entire ensemble of models.
torsion angles cluster together across the entire ensemble of models.
It is defined as
![]()
where
![]() ,
n being the number of members in the ensemble and
R is given by the expression
,
n being the number of members in the ensemble and
R is given by the expression
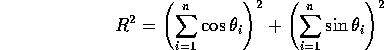
The value of the circular variance varies from 0 to 1, with the lower the value the tighter the clustering of the values about a single mean value.
For two dimensional distributions, such as the distributions of the
![]() -
-![]() values on the residue-by-residue Ramachandran plots, the
expression for
values on the residue-by-residue Ramachandran plots, the
expression for
![]() above is modified to
above is modified to

Allen, F. H. & Johnson, O. (1991) Automated conformational analysis from crystallographic data. 4. Statistical descriptors for a distribution of torsion angles. Acta Cryst., B47, 62-67.
MacArthur M W & Thornton J M (1993). Conformational analysis of protein structures derived from NMR data. Proteins, 17, 232-251.