




 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots





 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots
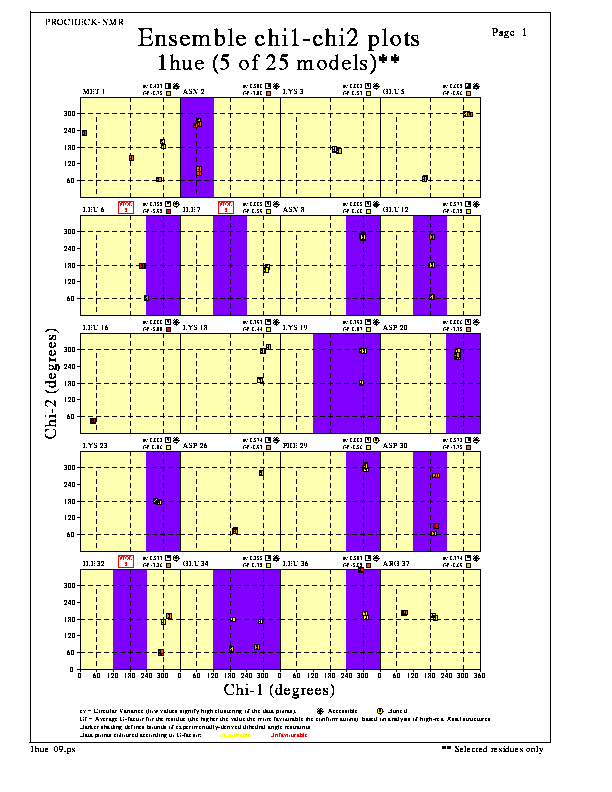
The Residue chi1-chi2 plots show how each residue's chi1-chi2 dihedral angles vary for each residue across the different models in the NMR ensemble. The corresponding model number is shown inside each data-point. If you want to know where on the plot each of the models lands, you can select the splay-numbering option in the procheck_nmr.prm parameter file.
The colours of the data points vary according to how favourable (yellow) or unfavourable (red) the chi1-chi2 values are for the given residue type.
The dotted lines correspond to the most-favoured chi-1 and chi-2 conformations: namely, gauche minus, trans and chi1 gauche. One would normally expect the values to cluster around the positions where these dotted lines cross; however, for some residues - most notably Asn, Asp and Trp - the most favourable chi-2 conformations tend to be displaced slightly from the above-named conformations.
Where any of the residues have had restraints applied to the chi-1 and/or chi-2 dihedral angles, the ranges of these are shown on the plots as the darker bands or squares. The numbers of violations of these restraints, if any, are printed on the plot.
It is sometimes possible to see how the values cluster at the borders of these restraint ranges, as though trying to escape from the range, suggesting that maybe the range itself may be incorrect.
Above the top right-hand corner of each residue's graph is shown:-
The main options for the Residue chi1-chi2 plots are:-
These options can be altered by editing the parameter file, procheck_nmr.prm, as described here.





 PROCHECK-NMR - Sample plots
PROCHECK-NMR - Sample plots