Glucan 1,3-beta-glucosidase
Fungal exo-beta-(1,3)-glucanases, such as that from the human pathogen Candida albicans, belong to the glycosyl hydrolase family 5 that also includes many bacterial cellulases. The Candida albicans enzyme is involved in the metabolism of cell wall glucan. It catalyses the hydrolytic removal of glucose residues from the non-reducing end of beta-1,3-glucan and (to a lesser extent) beta-1,6-glucan, the two main structural components of the Candida albicans cell wall. The enzyme also has glucosyl transferase activity in which a molecule other than water can accept the removed glucose residue. This may be important in shaping the cell wall during morphogenesis.
Reference Protein and Structure
- Sequence
-
P29717
 (2.4.1.-, 3.2.1.58)
(2.4.1.-, 3.2.1.58)
 (Sequence Homologues)
(PDB Homologues)
(Sequence Homologues)
(PDB Homologues)
- Biological species
-
Candida albicans SC5314 (Yeast)

- PDB
-
1cz1
- EXO-B-(1,3)-GLUCANASE FROM CANDIDA ALBICANS AT 1.85 A RESOLUTION
(1.85 Å)



- Catalytic CATH Domains
-
3.20.20.80
 (see all for 1cz1)
(see all for 1cz1)
Enzyme Reaction (EC:3.2.1.58)
Enzyme Mechanism
Introduction
The reaction proceeds by a double displacement mechanism with net retention of configuration at the anomeric C1 position. Nucleophilic attack by Glu 292 leads to displacement of the leaving group and formation of a covalent glucosyl-enzyme intermediate via a presumed oxo-carbenium ion-like transition state. Glu 192 acts as a general acid to protonate the departing leaving group.
Catalytic Residues Roles
| UniProt | PDB* (1cz1) | ||
| Glu230 | Glu192(186)A | Acts as a general acid to protonate the leaving group in formation of the glycosyl-enzyme intermediate. Later acts as a general base to abstract a proton from water so that it can carry out a nucleophilic attack on the the intermediate. | proton acceptor, proton donor, activator, increase nucleophilicity, promote heterolysis |
| Glu330 | Glu292(286)A | Attacks the glycosidic bond to form a glycosyl-enzyme intermediate via a presumed oxo-carbenium ion-like transition state. | covalently attached, nucleofuge, nucleophile |
Chemical Components
overall product formed, overall reactant used, intermediate formation, bimolecular nucleophilic substitution, proton transfer, hydrolysis, intermediate terminatedReferences
- Cutfield SM et al. (1999), J Mol Biol, 294, 771-783. The structure of the exo-β-(1,3)-glucanase from Candida albicans in native and bound forms: relationship between a pocket and groove in family 5 glycosyl hydrolases. DOI:10.1006/jmbi.1999.3287. PMID:10610795.
- Mackenzie LF et al. (1997), J Biol Chem, 272, 3161-3167. Identification of Glu-330 as the Catalytic Nucleophile of Candida albicans Exo- -(1,3)-glucanase. DOI:10.1074/jbc.272.6.3161. PMID:9013549.
- Chambers RS et al. (1993), FEBS Lett, 327, 366-369. Identification of a putative active site residue in the exo-β-(1,3)-glucanase ofCandida albicans. DOI:10.1016/0014-5793(93)81022-r. PMID:8348966.
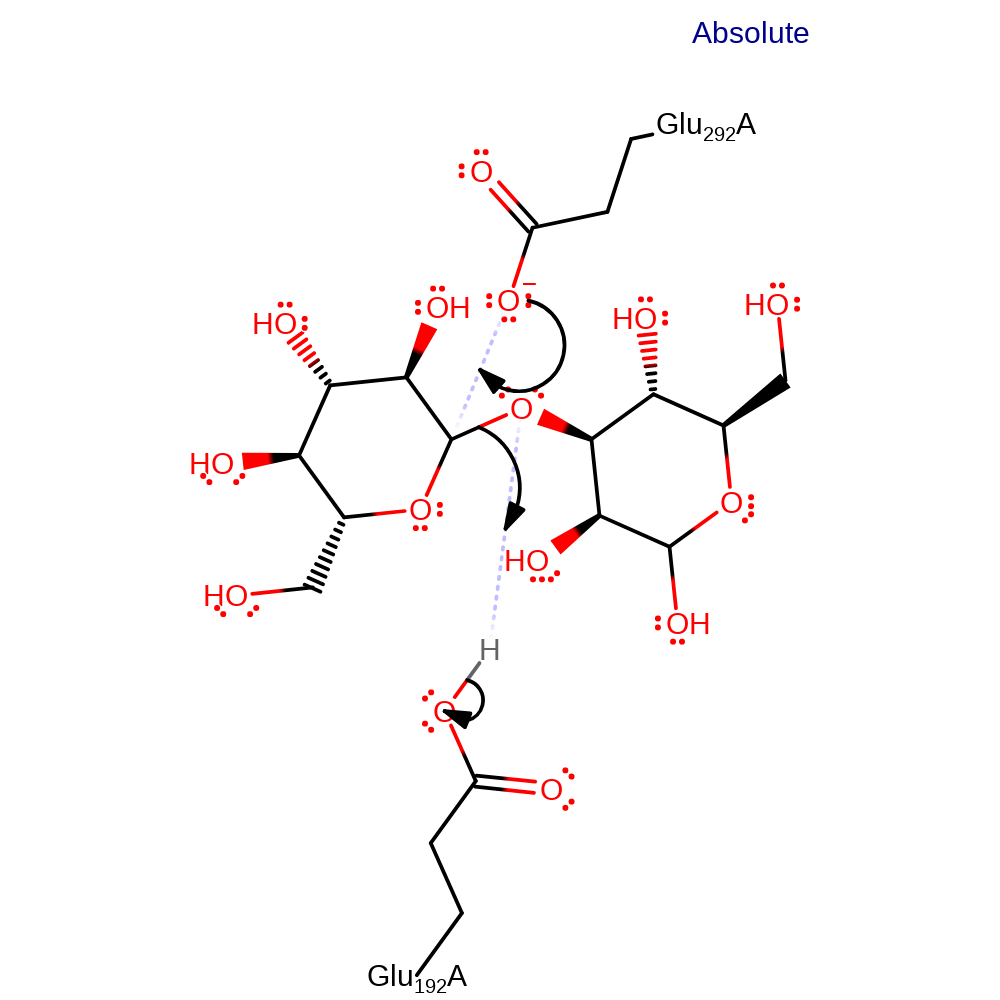
Step 1. Glu292 performs a nucleophilic attack on C1, this leads to the displacement of the leaving group which is protonated by Glu192.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Glu292(286)A | covalently attached |
| Glu192(186)A | promote heterolysis |
| Glu292(286)A | nucleophile |
| Glu192(186)A | proton donor |
Chemical Components
overall product formed, overall reactant used, intermediate formation, ingold: bimolecular nucleophilic substitution, proton transfer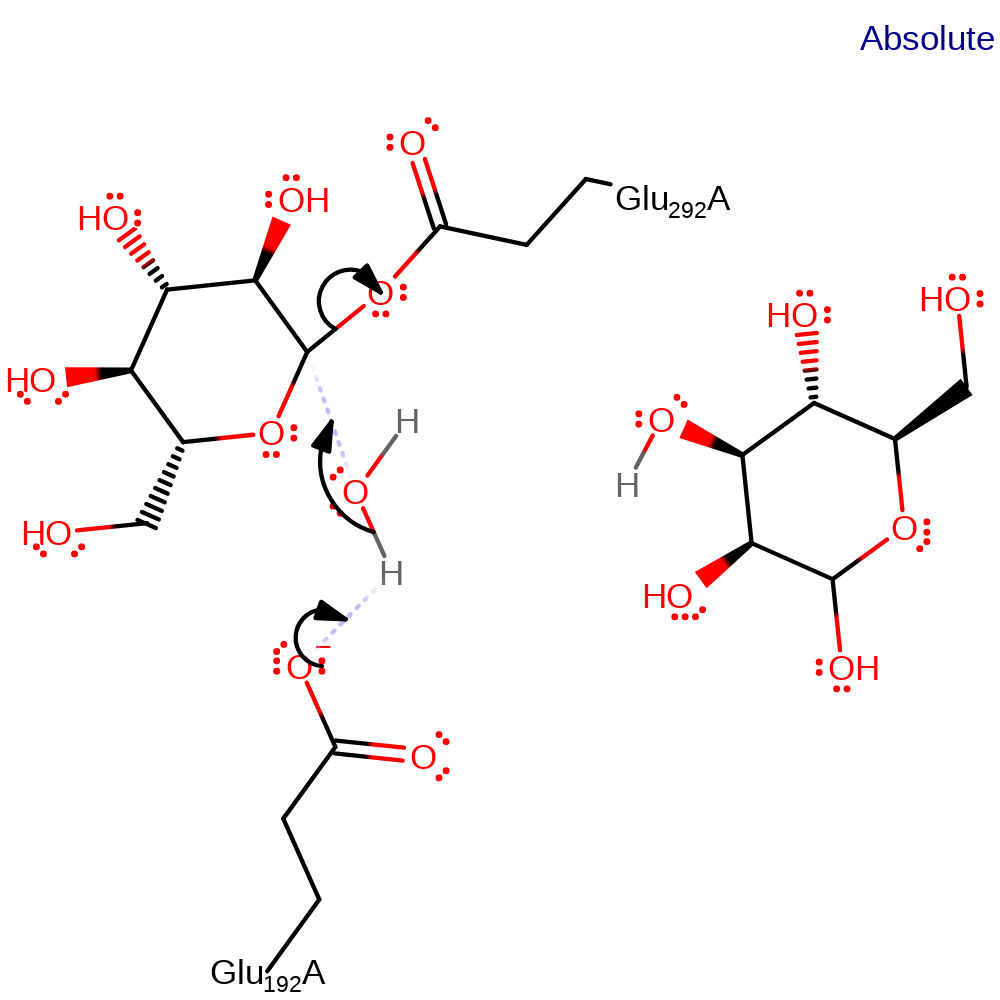
Step 2. Glu192 activates a water molecule which hydrolyses the intermediate.
Download: Image, Marvin FileCatalytic Residues Roles
| Residue | Roles |
|---|---|
| Glu192(186)A | activator, increase nucleophilicity, proton acceptor |
| Glu292(286)A | nucleofuge |



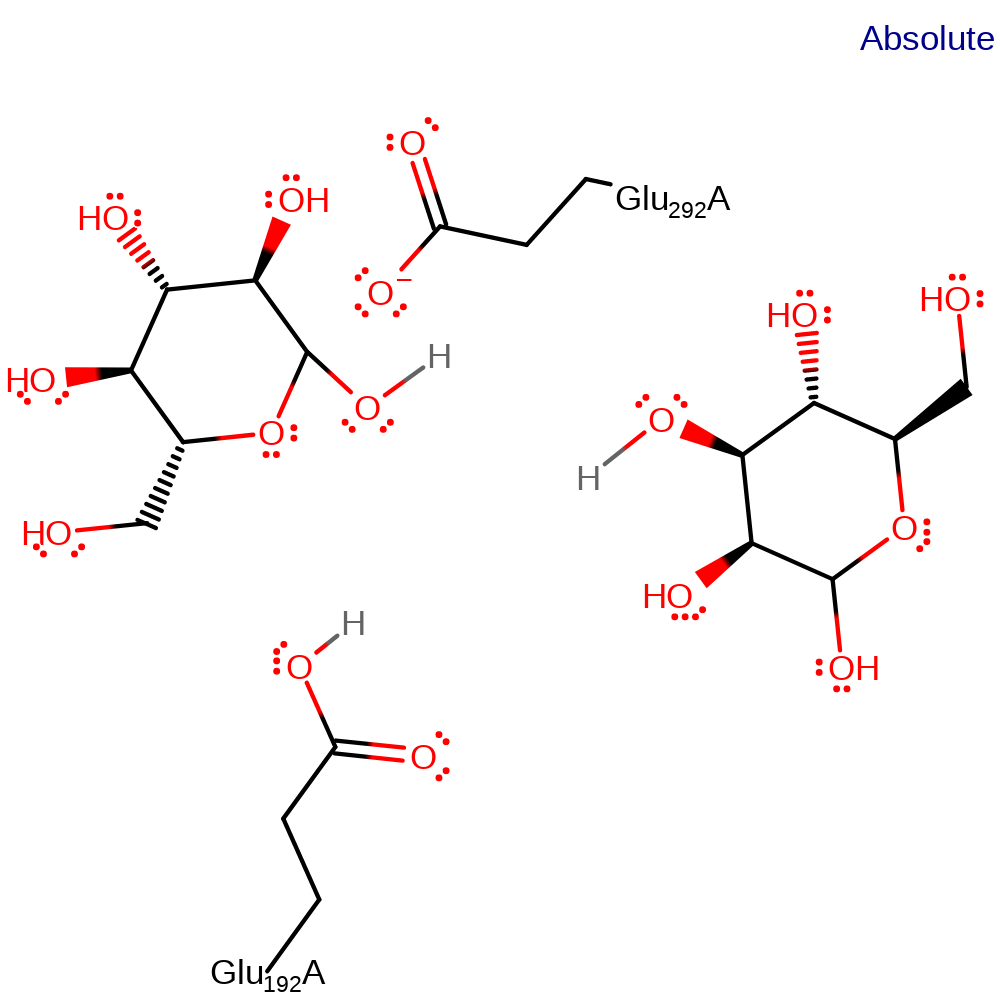 Download:
Download: