 |
PDBsum entry 2fxe

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.49
- RNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
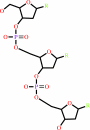 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.3.1.13.2
- exoribonuclease H.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Exonucleolytic cleavage to 5'-phosphomonoester oligonucleotides in both 5'- to 3'- and 3'- to 5'-directions.
|
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.3.1.26.13
- retroviral ribonuclease H.
|
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.3.4.23.16
- HIV-1 retropepsin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Specific for a P1 residue that is hydrophobic, and P1' variable, but often Pro.
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
J Virol
81:9525-9535
(2007)
|
|
PubMed id:
|
|
|
|

|
| |
|
X-ray crystal structures of human immunodeficiency virus type 1 protease mutants complexed with atazanavir.
|
|
H.E.Klei,
K.Kish,
P.F.Lin,
Q.Guo,
J.Friborg,
R.E.Rose,
Y.Zhang,
V.Goldfarb,
D.R.Langley,
M.Wittekind,
S.Sheriff.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Atazanavir, which is marketed as REYATAZ, is the first human immunodeficiency
virus type 1 (HIV-1) protease inhibitor approved for once-daily administration.
As previously reported, atazanavir offers improved inhibitory profiles against
several common variants of HIV-1 protease over those of the other peptidomimetic
inhibitors currently on the market. This work describes the X-ray crystal
structures of complexes of atazanavir with two HIV-1 protease variants, namely,
(i) an enzyme optimized for resistance to autolysis and oxidation, referred to
as the cleavage-resistant mutant (CRM); and (ii) the M46I/V82F/I84V/L90M mutant
of the CRM enzyme, which is resistant to all approved HIV-1 protease inhibitors,
referred to as the inhibitor-resistant mutant. In these two complexes,
atazanavir adopts distinct bound conformations in response to the V82F
substitution, which may explain why this substitution, at least in isolation,
has yet to be selected in vitro or in the clinic. Because of its nearly
symmetrical chemical structure, atazanavir is able to make several analogous
contacts with each monomer of the biological dimer.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
S.K.Law,
R.R.Wang,
A.N.Mak,
K.B.Wong,
Y.T.Zheng,
and
P.C.Shaw
(2010).
A switch-on mechanism to activate maize ribosome-inactivating protein for targeting HIV-infected cells.
|
| |
Nucleic Acids Res,
38,
6803-6812.
|
 |
|
|
|
|
 |
J.M.Sayer,
F.Liu,
R.Ishima,
I.T.Weber,
and
J.M.Louis
(2008).
Effect of the active site D25N mutation on the structure, stability, and ligand binding of the mature HIV-1 protease.
|
| |
J Biol Chem,
283,
13459-13470.
|
 |
|
PDB codes:
|
 |
|
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

