 |
PDBsum entry 7lec

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Hydrolase/hydrolase inhibitor
|
PDB id
|
|

|

|
7lec
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Hydrolase/hydrolase inhibitor
|
 |
|
Title:
|
 |
HIV-1 protease wt (nl4-3) in complex with pu3 (lr3-69)
|
|
Structure:
|
 |
Protease. Chain: b, a. Engineered: yes
|
|
Source:
|
 |
Human immunodeficiency virus 1. Organism_taxid: 11676. Gene: pol. Expressed in: escherichia coli. Expression_system_taxid: 562.
|
|
Resolution:
|
 |
|
1.94Å
|
R-factor:
|
0.202
|
R-free:
|
0.247
|
|
|
Authors:
|
 |
G.J.Lockbaum,L.N.Rusere,M.Henes,K.Kosovrasti,S.K.Lee,E.Spielvogel, E.A.Nalivaika,R.Swanstrom,N.Kurtyilmaz,C.A.Schiffer,A.Ali
|
|
Key ref:
|
 |
G.J.Lockbaum
et al.
Hiv-1 protease inhibitors with a p1 phosphonate modif maintain potency against drug resistant variants by increased van der waals contacts with flaps residues.
To be published,
.
|
 |
|
Date:
|
 |
|
14-Jan-21
|
Release date:
|
26-Jan-22
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P12497
(POL_HV1N5) -
Gag-Pol polyprotein from Human immunodeficiency virus type 1 group M subtype B (isolate NY5)
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
1435 a.a.
99 a.a.*
|

|

|

|

|
|

|

|

|
 |
 |

|
|
Key: |
 |
PfamA domain |
 |
 |
 |
Secondary structure |
 |
|
*
PDB and UniProt seqs differ
at 1 residue position (black
cross)
|
|

|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.7.7.49
- RNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
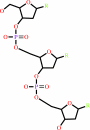 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.3.1.-.-
|
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.3.1.13.2
- exoribonuclease H.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Exonucleolytic cleavage to 5'-phosphomonoester oligonucleotides in both 5'- to 3'- and 3'- to 5'-directions.
|
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.3.1.26.13
- retroviral ribonuclease H.
|
|
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.4.23.16
- HIV-1 retropepsin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Specific for a P1 residue that is hydrophobic, and P1' variable, but often Pro.
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |

