 |
PDBsum entry 7h2c

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Viral protein
|
PDB id
|
|

|

|
7h2c
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Viral protein
|
 |
|
Title:
|
 |
Pandda analysis group deposition -- crystal structure of zikv ns2b-ns3 protease in complex with z1328968520
|
|
Structure:
|
 |
Serine protease subunit ns2b. Chain: a. Synonym: flavivirin protease ns2b regulatory subunit,non-structural protein 2b. Engineered: yes. Serine protease ns3. Chain: b. Synonym: flavivirin protease ns3 catalytic subunit,non-structural protein 3.
|
|
Source:
|
 |
Zika virus. Organism_taxid: 64320. Expressed in: escherichia coli. Expression_system_taxid: 562. Expression_system_taxid: 562
|
|
Resolution:
|
 |
|
1.40Å
|
R-factor:
|
0.223
|
R-free:
|
0.249
|
|
|
Authors:
|
 |
X.Ni,A.S.Godoy,P.G.Marples,M.Fairhead,B.H.Balcomb,C.W.E.Tomlinson, L.Koekemoer,J.C.Aschenbrenner,R.M.Lithgo,W.Thompson,C.Wild, E.P.Williams,M.Winokan,A.V.Chandran,D.Fearon,M.A.Walsh,F.Von Delft
|
|
Key ref:
|
 |
X.Ni
et al.
Pandda analysis group deposition of zikv ns2b-Ns3 pro.
To be published,
.
|
 |
|
Date:
|
 |
|
03-Apr-24
|
Release date:
|
08-May-24
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
Chains A, B:
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
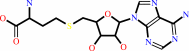 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
Chains A, B:
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
Chains A, B:
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
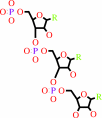 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
Chains A, B:
E.C.3.4.21.91
- flavivirin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Selective hydrolysis of Xaa-Xaa-|-Xbb bonds in which each of the Xaa can be either Arg or Lys and Xbb can be either Ser or Ala.
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
Chains A, B:
E.C.3.6.1.15
- nucleoside-triphosphate phosphatase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a ribonucleoside 5'-triphosphate + H2O = a ribonucleoside 5'-diphosphate + phosphate + H+
|
 |
 |
 |
 |
 |
ribonucleoside 5'-triphosphate
|
+
|
H2O
|
=
|
ribonucleoside 5'-diphosphate
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
Chains A, B:
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
|

