 |
PDBsum entry 6ecl
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
Chains A, B:
E.C.2.7.7.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
Chains A, B:
E.C.2.7.7.49
- RNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
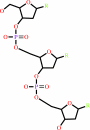 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
Chains A, B:
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
Chains A, B:
E.C.3.1.-.-
|
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
Chains A, B:
E.C.3.1.13.2
- exoribonuclease H.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Exonucleolytic cleavage to 5'-phosphomonoester oligonucleotides in both 5'- to 3'- and 3'- to 5'-directions.
|
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
Chains A, B:
E.C.3.1.26.13
- retroviral ribonuclease H.
|
|
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
Chains A, B:
E.C.3.4.23.16
- HIV-1 retropepsin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Specific for a P1 residue that is hydrophobic, and P1' variable, but often Pro.
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
|

