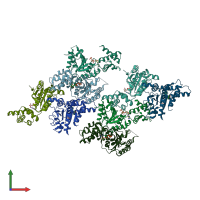
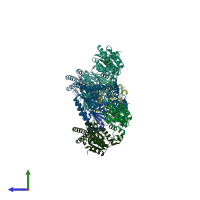
Assemblies
Multimeric state:
monomeric
Accessible surface area:
12621.64 Å2
Buried surface area:
778.64 Å2
Dissociation area:
389.32
Å2
Dissociation energy (ΔGdiss):
4.55
kcal/mol
Dissociation entropy (TΔSdiss):
5.33
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12315.89 Å2
Buried surface area:
765.97 Å2
Dissociation area:
382.99
Å2
Dissociation energy (ΔGdiss):
3.81
kcal/mol
Dissociation entropy (TΔSdiss):
5.3
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12413.73 Å2
Buried surface area:
767.6 Å2
Dissociation area:
383.8
Å2
Dissociation energy (ΔGdiss):
4.98
kcal/mol
Dissociation entropy (TΔSdiss):
5.31
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12282.71 Å2
Buried surface area:
766.69 Å2
Dissociation area:
383.34
Å2
Dissociation energy (ΔGdiss):
4.42
kcal/mol
Dissociation entropy (TΔSdiss):
5.32
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12608.07 Å2
Buried surface area:
762.93 Å2
Dissociation area:
381.47
Å2
Dissociation energy (ΔGdiss):
4.1
kcal/mol
Dissociation entropy (TΔSdiss):
5.31
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12316.8 Å2
Buried surface area:
773.85 Å2
Dissociation area:
386.92
Å2
Dissociation energy (ΔGdiss):
4.98
kcal/mol
Dissociation entropy (TΔSdiss):
5.32
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12453.27 Å2
Buried surface area:
775.07 Å2
Dissociation area:
387.54
Å2
Dissociation energy (ΔGdiss):
4.87
kcal/mol
Dissociation entropy (TΔSdiss):
5.32
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Multimeric state:
monomeric
Accessible surface area:
12344.38 Å2
Buried surface area:
767.19 Å2
Dissociation area:
383.6
Å2
Dissociation energy (ΔGdiss):
5.25
kcal/mol
Dissociation entropy (TΔSdiss):
5.31
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-152572
Macromolecules
Chains: A, B, C, D, E, F, G, H
Length: 270 amino acids
Theoretical weight: 30.02 KDa
Source organisms: Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 270 amino acids
Theoretical weight: 30.02 KDa
Source organisms: Expression system: Escherichia coli
UniProt:
- Canonical:
 Q8U4H3 (Residues: 125-309; Coverage: 47%)
Q8U4H3 (Residues: 125-309; Coverage: 47%) - Canonical:
 P33297 (Residues: 356-434; Coverage: 18%)
nullnull
P33297 (Residues: 356-434; Coverage: 18%)
nullnull
Pfam:
InterPro:
- P-loop containing nucleoside triphosphate hydrolase
- AAA+ ATPase domain
- ATPase, AAA-type, core
- ATPase, AAA-type, conserved site
- AAA ATPase, AAA+ lid domain