Assemblies
Assembly Name:
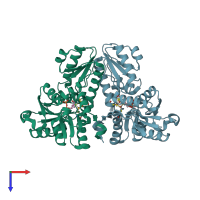
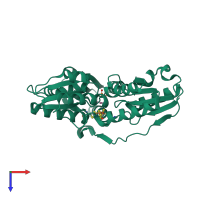
4-hydroxy-3-methylbut-2-enyl diphosphate reductase
Multimeric state:
monomeric
Accessible surface area:
14251.32 Å2
Buried surface area:
374.07 Å2
Dissociation area:
187.04
Å2
Dissociation energy (ΔGdiss):
16.1
kcal/mol
Dissociation entropy (TΔSdiss):
3.19
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-158657
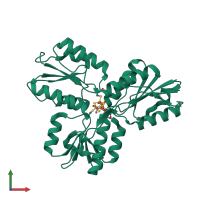
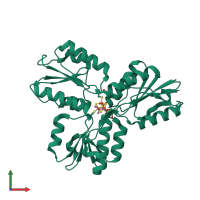
Assembly Name:
4-hydroxy-3-methylbut-2-enyl diphosphate reductase
Multimeric state:
monomeric
Accessible surface area:
14459.14 Å2
Buried surface area:
374.66 Å2
Dissociation area:
187.33
Å2
Dissociation energy (ΔGdiss):
16.13
kcal/mol
Dissociation entropy (TΔSdiss):
3.19
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-158657
Macromolecules
Chains: A, B
Length: 328 amino acids
Theoretical weight: 36.22 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
Pfam: LytB protein
InterPro: 4-hydroxy-3-methylbut-2-enyl diphosphate reductase
CATH:
Length: 328 amino acids
Theoretical weight: 36.22 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P62623 (Residues: 1-316; Coverage: 100%)
P62623 (Residues: 1-316; Coverage: 100%)
Pfam: LytB protein
InterPro: 4-hydroxy-3-methylbut-2-enyl diphosphate reductase
CATH: