Assemblies
Assembly Name:
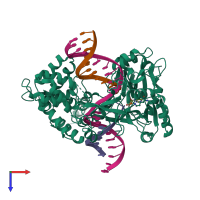
DNA ligase 3 and DNA
Multimeric state:
hetero tetramer
Accessible surface area:
29878.73 Å2
Buried surface area:
7539.43 Å2
Dissociation area:
1,162.54
Å2
Dissociation energy (ΔGdiss):
21.42
kcal/mol
Dissociation entropy (TΔSdiss):
10.16
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-116515
Macromolecules
Chain: A
Length: 579 amino acids
Theoretical weight: 65.68 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 579 amino acids
Theoretical weight: 65.68 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P49916 (Residues: 257-833; Coverage: 57%)
P49916 (Residues: 257-833; Coverage: 57%)
Pfam:
InterPro:
- DNA ligase, ATP-dependent, N-terminal domain superfamily
- DNA ligase, ATP-dependent, N-terminal
- DNA ligase, ATP-dependent
- DNA ligase, ATP-dependent, central
- DNA ligase, ATP-dependent, conserved site
- Nucleic acid-binding, OB-fold
- DNA ligase, ATP-dependent, C-terminal
Name:
5'-D(P*CP*GP*GP*GP*AP*TP*GP*CP*GP*TP*C)-3'
Representative chains: B
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.39 KDa
Representative chains: B
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.39 KDa
Name:
5'-D(*GP*TP*CP*GP*GP*AP*CP*TP*G)-3'
Representative chains: C
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 9 nucleotides
Theoretical weight: 2.77 KDa
Representative chains: C
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 9 nucleotides
Theoretical weight: 2.77 KDa
Name:
5'-D(*GP*CP*CP*AP*GP*TP*CP*CP*GP*AP*CP*GP*AP*CP*GP*CP*AP*TP*CP*CP*CP*G)-3'
Representative chains: D
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 22 nucleotides
Theoretical weight: 6.68 KDa
Representative chains: D
Source organism: Homo sapiens [9606]
Expression system: Not provided
Length: 22 nucleotides
Theoretical weight: 6.68 KDa