 |
PDBsum entry 4kld

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase, lyase/DNA
|
PDB id
|
|

|

|
4kld
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
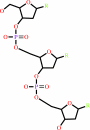 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.4.2.99.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.4.2.99.18
- DNA-(apurinic or apyrimidinic site) lyase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
2'-deoxyribonucleotide-(2'-deoxyribose 5'-phosphate)- 2'-deoxyribonucleotide-DNA = a 3'-end 2'-deoxyribonucleotide-(2,3- dehydro-2,3-deoxyribose 5'-phosphate)-DNA + a 5'-end 5'-phospho- 2'-deoxyribonucleoside-DNA + H+
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Cell
154:157-168
(2013)
|
|
PubMed id:
|
|
|
|

|
| |
|
Observing a DNA polymerase choose right from wrong.
|
|
B.D.Freudenthal,
W.A.Beard,
D.D.Shock,
S.H.Wilson.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
DNA polymerase (pol) β is a model polymerase involved in gap-filling DNA
synthesis utilizing two metals to facilitate nucleotidyl transfer. Previous
structural studies have trapped catalytic intermediates by utilizing substrate
analogs (dideoxy-terminated primer or nonhydrolysable incoming nucleotide). To
identify additional intermediates during catalysis, we now employ natural
substrates (correct and incorrect nucleotides) and follow product formation in
real time with 15 different crystal structures. We are able to observe molecular
adjustments at the active site that hasten correct nucleotide insertion and
deter incorrect insertion not appreciated previously. A third metal binding site
is transiently formed during correct, but not incorrect, nucleotide insertion.
Additionally, long incubations indicate that pyrophosphate more easily
dissociates after incorrect, compared to correct, nucleotide insertion. This
appears to be coupled to subdomain repositioning that is required for catalytic
activation/deactivation. The structures provide insights into a fundamental
chemical reaction that impacts polymerase fidelity and genome stability.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

