
The work of bees in pollinating our crops is as vital now as it has been for centuries, but bees are having a hard time at the moment for a variety of reasons, including viral diseases. Recent structures of a bee virus show novel features that might contribute to its infectivity.
Slow Bee Paralysis Virus (SBPV, shown left along with a honey bee) is one of several viruses in the Iflaviridae family which can infect honey bees. Two crystal structures of this RNA virus (PDB entries 5j96 and 5j98) showed it to have the icosahedral capsid formed from sixty copies of three proteins which is common to picornaviruses. However one of these proteins, VP3, contains an extra domain termed the P-domain (coloured orange in the image to the left). This has not been seen before in such viruses, although it does occur in some viruses which infect plants. The P-domain (P for protruding) forms ‘crowns’ on the surface of the virus but the position of the crown differs between the two crystal structures, suggesting a degree of mobility of this domain.
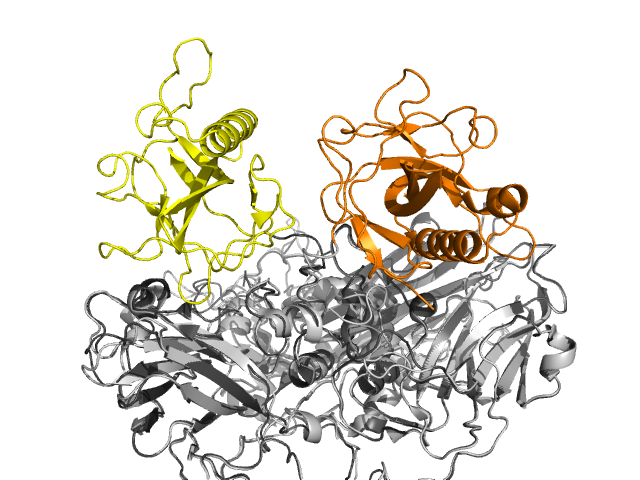
Superimposition of both structures of SBPV. Only the three-protein building block is shown. The P-domains are coloured yellow and orange, showing how they differ between the two forms.
As the domain is surface-exposed it is in a position to encounter and bind molecules on host cells within the bee, so may act as a receptor. The domain contains a serine, histidine and aspartate residue positioned almost as a catalytic triad, a very familiar motif in many hydrolytic enzymes such as chymotrypsin. Whilst not optimally positioned for catalysis, some movement may occur on binding to a bee cell-surface which activates the enzyme, but more study is needed to discover if this is the case and, if so, the identity of the substrate.
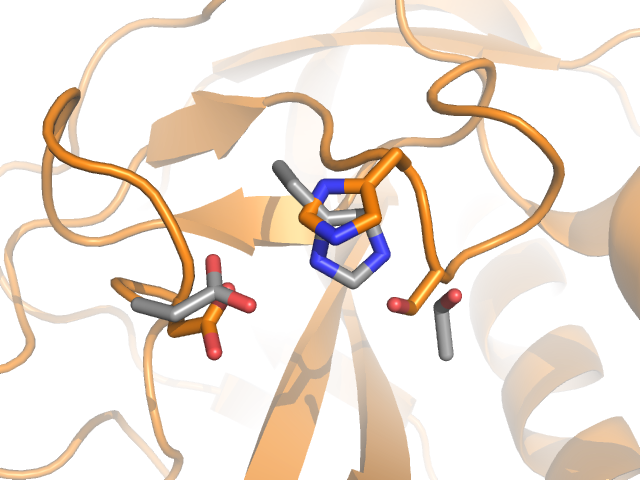
Detail of the P-domain with the putative catalytic triad coloured orange and shown as sticks. The catalytic triad of chymotrypsin (PDB entry 1yph) is overlaid and shown as grey sticks.
The work by Kalynych et al is published in the Journal of Virology, doi:10.1128/JVI.00680-16


