What do we do?
The eQTL Catalogue aims to provide uniformly processed gene expression and splicing QTLs from all available public studies on human.
Quantitative trait loci (QTL) are genomic variants that are significantly associated to a measurable phenotype. This resource focuses on expression cis-QTLs (cis-eQTL) where variants are associated to expression levels of nearby genes, and on splicing QTLs (sQTL) where variants are associated to specific splicing events on nearby splice junctions.
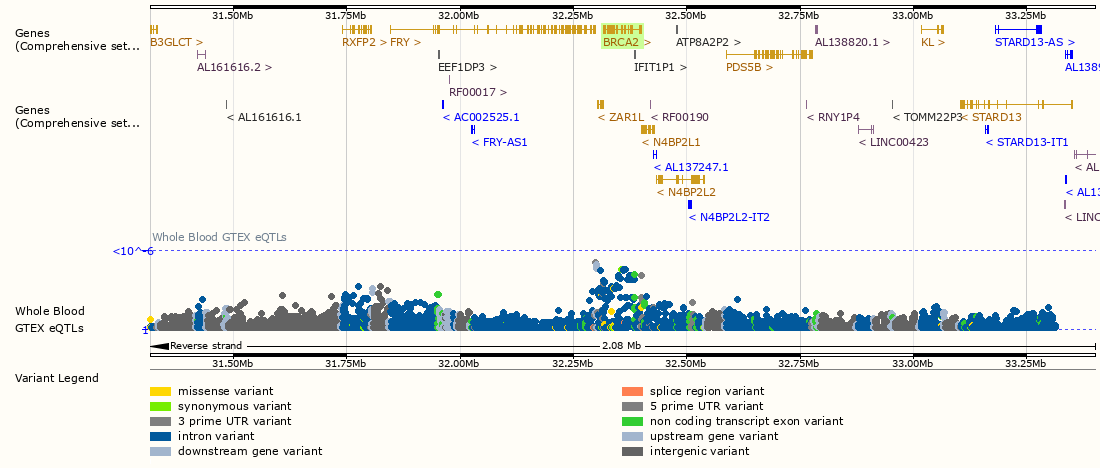
They have proven particularly useful to (statistical) geneticists exploring GWAS results, who wish to associate non-coding GWAS SNP associations to a molecular mechanism, such as perturbed gene expression or splicing.
To compute them we process public studies with our uniform pipeline. We currently provide access to 32 published studies, with an aim to cover all accessible studies on human.
A complete explanation can be found in our flagship paper and a presentation on our webinar.
Access our results
You can access our results via FTP in CSV and HDF5 formats, or query part of our data via our RESTful API. Our results are also displayed on the Open Targets Platform. More information on our re-use policy is on the License tab.
Let us know what you think!
Feel free to send us your feedback at eqtlcatalogue@ebi.ac.uk
Follow us
Keep up to date on the eQTL Catalogue through announcements on our Twitter account @eQTLCatalogue.