EMPIAR deposition system
EMPIAR deposition manual
Contents
1 Introduction
EMPIAR, the Electron Microscopy Public Image Archive, is a public resource for raw images underpinning 3D cryo-EM maps and tomograms (themselves archived in EMDB). EMPIAR also accommodates 3D datasets obtained with volume EM techniques and soft and hard X-ray tomography. The EMPIAR Deposition System can be used to deposited data to EMPIAR. All EMPIAR entries (with certain exceptions, see below) are required to be associated with one or more EMDB entries. “Associated” in this context means that it should be the image data used to obtain the 3D reconstruction(s) deposited as one or more EMDB entries. In such cases depositors are encouraged to inform EMDB and PDB (as appropriate) about the EMPIAR accession.
EMPIAR will accept data that is not associated with an EMDB entry in the following cases:
- 2D/3D data from 3D imaging modalities not covered by EMDB (e.g. 3DSEM and SXT);
- 2D EM data used in integrative/hybrid methods, associated with a structure deposited in the PDB or PDB-Dev archive;
- Certain reference and benchmark datasets (to be decided on a case-by-case basis)*
- Datasets used for certain community challenges (such as the 2015 Map Validation Challenge, see: “The first single particle analysis Map Challenge: A summary of the assessments,” J. Struct. Biol. 204 (2018), 291-300, https://doi.org/10.1016/j.jsb.2018.08.010)*
* We are keen to support community challenges and archival of reference data sets. Please contact the operators of the EMPIAR archive prior to deposition.
In cases not covered above, please contact the operators of the EMPIAR archive prior to deposition to discuss the potential suitability of EMPIAR for your data.
2 Pre-deposition preparations
In order to make the deposition process run smoothly we request that you make certain preparations prior to deposition:
- If, as described above, you need to - make a note of the accession code (in the form EMD-####, e.g., EMD-1001) of the related EMDB deposition. Not only is this a requirement, but the accession code can also be used to automatically fill in fields (if the EMDB entry has been released).
- Organize your data
- If you are planning to upload multiple datasets (e.g., micrographs and particle stacks) we highly recommend that you create one sub-directory for each dataset.
- Please name your subdirectories so that it is easy to understand the organization. For example "micrographs" for micrographs and "particles" for particles.
- Typically having more than 4000 files in a directory has a tendency to slow down access considerably. We would recommend in this case that you sub-divide the directory into subdirectories with no more than 4000 files each.
- If you have a single file larger than 1 TB please contact us in advance.
- Make a note of the details describing each dataset that you will be asked for during the deposition
process. These include:
- Are the images processed or raw?
- Are they multi-frame images? If so which frames have been used?
- Image format
- Number of images (or tilt series)
- Image width and height
- Pixel size
- Pixel type (unsigned byte, byte, 32-bit float etc). EMPIAR system supports automated
reading of headers of many common formats such as MRC, TIFF, DM4, etc., so you can
skip this step. You can also see this manually, for example,
by examining the header of the file with tools such as IMOD, BSOFT or EMAN2 or, as
suggested by one of our users, Takanori Nakane, by using tiffdump command:
tiffdump -h filename.tif
3 Data transfer technologies
In order to upload data to EMPIAR we provide two alternatives which are both capable of dealing efficiently and robustly with the large data volumes associated with EMPIAR — Globus (https://www.globus.org) and Aspera (https://www.ibm.com/aspera/connect/). Both technologies require you to install some software on your machine but are free at the point of use.
3.1 Globus
Please follow the official guide to register (this is free) and set up Globus.
Once you have set up your local endpoint, you can try downloading from EMPIAR. To do so in Collection search enter "EMBL-EBI Public Data" Collection and set the directory to: /gridftp/empiar/world_availability/, select and activate the transfer as per the above mentioned guide.
3.2 Aspera
As easy way to check and install the Aspera plugins is to go to the EMPIAR website and try downloading an entry, Figure 1.

If Aspera is not installed it will prompt and guide you to install the relevant software, Figure 2.
Now clicking the download button will initiate the transfer (you can cancel the download once the transfer has started, Figure 3).

Aspera makes use of UDP transfer technology. Some institutes block the UDP port (port 33001) by default and it is not possible to get them enabled. If this is the case for you then we recommend that you use Globus which relies on GridFTP.
4 User accounts and deposition landing page
You need a user account to use the deposition system; to register please proceed to the registration page, Figure 4.
Once logged in, the "Edit profile" option in the left menu allows you to update your profile and change your password, Figure 5.

We recommend that you keep your profile up-to-date as you can use it to automatically fill in form fields in a deposition.
By default, when you log in you will be taken to the landing page, Figure 6, which presents the option to create new depositions ("Create a new deposition") and a table with depositions that you can access.

One user can create multiple depositions and multiple users can share access to one deposition. For any deposition, only one user is considered the "owner". The owner can grant access rights for the deposition to other users ("View only", "View and edit" or "view, edit and submit") or can transfer ownership to another user but following the "Change ownership/grant rights" link in the depositions table.
5 Automated deposition
It is possible to deposit into EMPIAR automatically using a Python script empiar-depositor. To do this you can use your credentials or generate a permanent token if you would prefer not to expose your EMPIAR credentials. The script takes as an input a JSON file with the description of your deposition. The JSON file corresponds to all the forms that you will be otherwise asked to fill in as described below. It can be automatically generated by Scipion or created manually according to the schema which can be found in the intallation location of the Python script or by the following link.
6 Manual deposition
6.1 Overview
The deposition process consists of three mandatory parts, Figure 7:
- providing the general metadata about the deposition — citation, title, authors, etc.;
- uploading the data — the transfer can take some time, so the next step most likely would not be undertaken immediately;
- associating the uploaded data with the corresponding image sets — that is, identifying the image sets present and describing them.

There is also an optional part where the depositor can provide segmentations of their data. This part can be activated from the image set page.
Once these steps have been completed, the deposition can be submitted. This will lock the deposition (make it uneditable) while it is being checked by the EMPIAR annotation team. They will communicate with the user regarding any issues and may choose to unlock the deposition if complementary details or data are required. Once this process is completed, the entry will be released to the public following the instructions provided (see below for options).
6.2 Form basics
6.2.1 Deposition locking
As multiple users can work on the same deposition and more than one user can have edit rights, a locking mechanism has been implemented to prevent simultaneous editing by multiple users.

Whenever you open a form page, the whole deposition becomes locked to you for 30 minutes and you have exclusive rights to edit it, Figure 8. It is possible to release the lock before the expiration time by closing all pages or by pressing the "Release Lock" button.
6.2.2 "Save", "Save & Validate", "Submit entry" and the traffic light system
Changes made to a form will be lost unless they are saved by pressing the "Save" or "Save & Validate" buttons. The former is for a temporary save of the page in case it is not possible to fill in all the mandatory information on the page in one go. However, to proceed with the submission it is necessary to have the information on the page validated by our system with "Save & Validate", Figure 9.

The state of the page is shown on the left-hand side menu, Figure 10. When the page is first opened, there is an empty circle next to its link, when it is saved the circle becomes filled with yellow, when it is validated with errors — red and when it successfully passes the validation — green.

When all forms have been filled out and validated successfully, a "Submit" button will become active. Please press "Submit" to send the deposition for review by the EMPIAR annotation team. The deposition will be locked from further editing from this stage onwards unless an annotator would require you to fill in or change any of the information.
6.2.3 Mandatory fields and the "N/A" button
Mandatory fields are marked in orange. You will also find many fields that have a "N/A" (not available/not applicable) button next to them, Figure 11. Not all of these fields are mandatory but we expect the user to at least press the "N/A" button to explicitly confirm that the information requested cannot be provided. Pressing the "N/A" will automatically erase the existing information in the form field and fill it with the special marker for N/A information.

6.2.4 Form field help and examples
Most form fields have a question mark symbol "?" next to them and an example value below them, Figure 12. Hovering over the question mark symbol will bring up a pop-up box with help.

6.3 Deposition overview page
6.3.1 Deposition image
This image will be used for representative purposes on the EMPIAR website alongside your entry, Figure 13. The image should be a minimum of 400 x 400 in png or gif format.
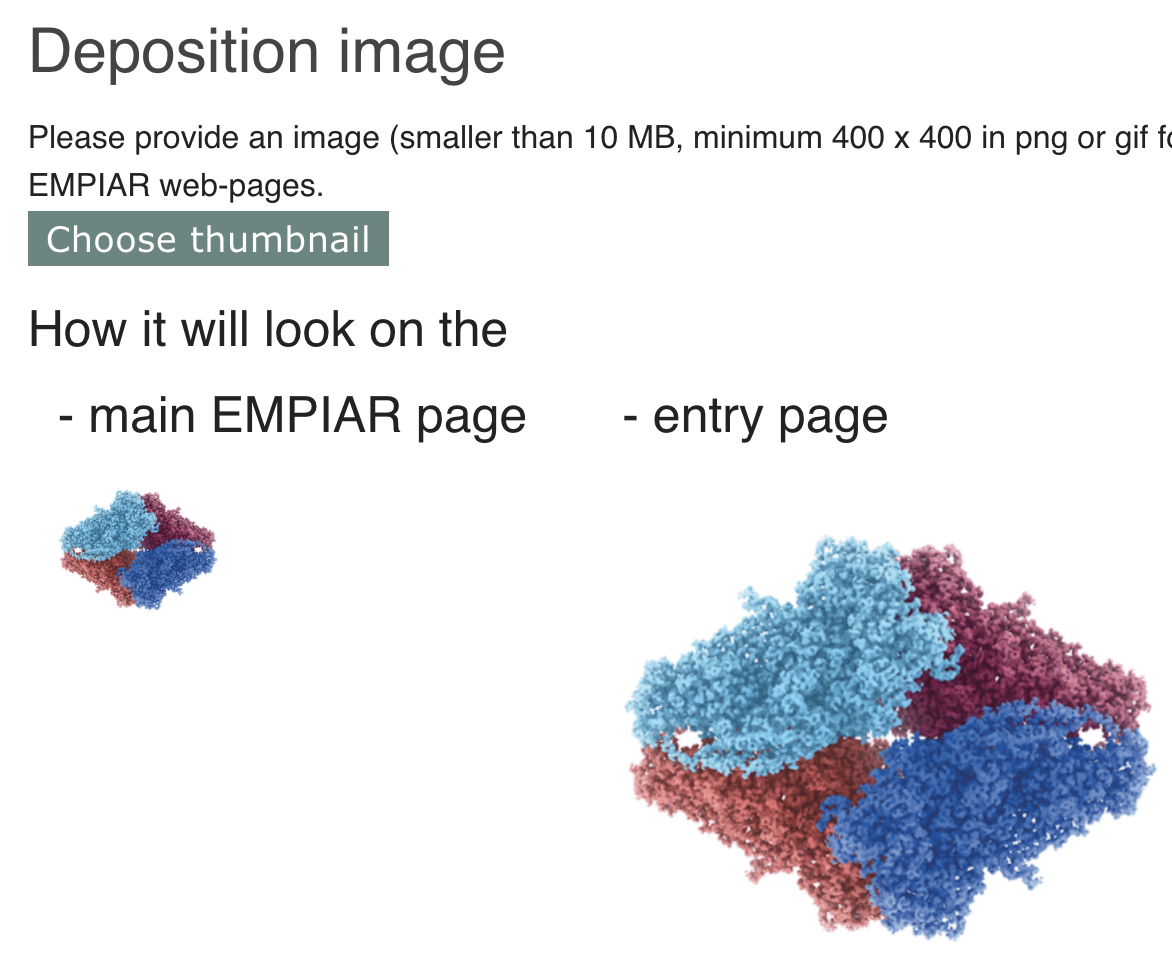

6.3.2 Harvesting information from the related EMDB entry and user profiles
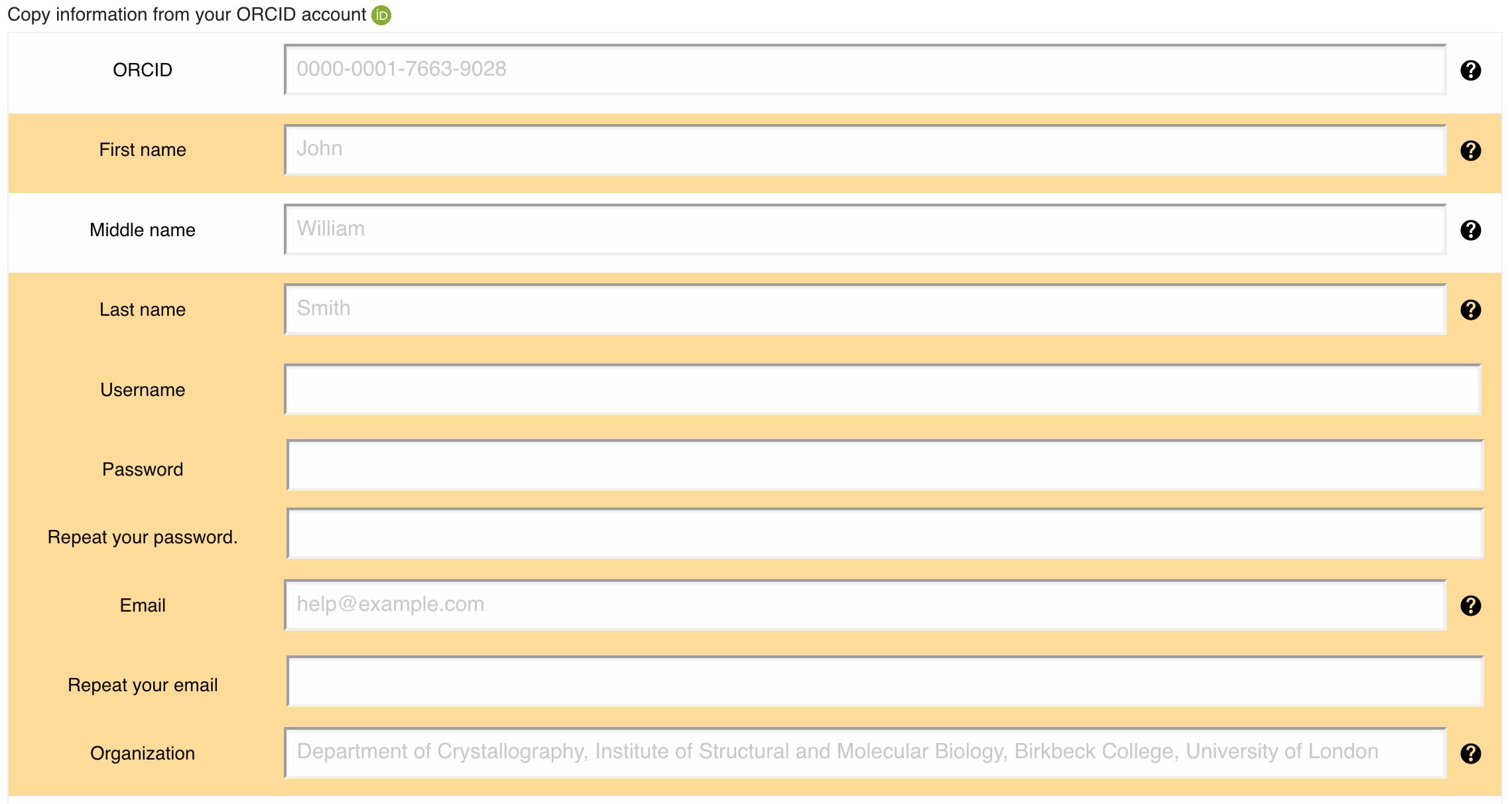
You can specify multiple EMDB accession codes but please note that there are separate boxes for released and unreleased entries, Figure 14. If the entry has been released you can copy authors from the related EMDB entry by pressing the "Fill in entry authors from the released EMDB entry" button.

You can copy authors from the related EMDB entry by filling in the "EMDB accession code" and then pressing the "Fill in entry authors from the released EMDB entry" button. You can also automatically populate the corresponding author and principal author fields from your public ORCID infromation or from the profile of any EMPIAR user that are associated with the deposition. Once the corresponding author fields have been populated, you may also copy these over to the principal author fields.
6.3.3 Citation information
Please provide the information about the citation related to your deposition, Figure 15.
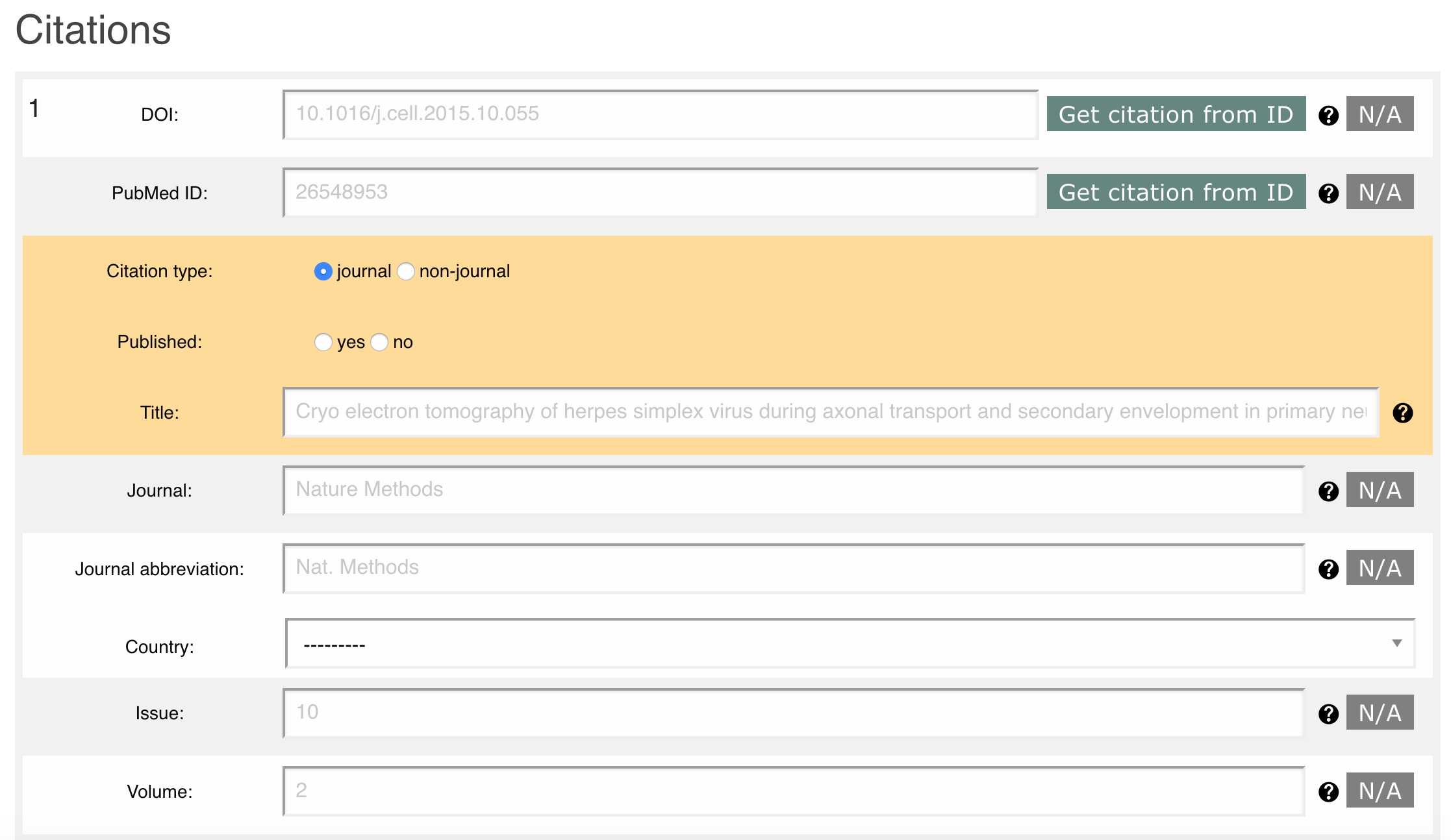
You can automatically fill in the citation information using DOI or PubMed ID.
If the information regarding editors is not available, please ignore the corresponding form.
6.3.4 Release instruction
EMPIAR depositions are released (made available to the public) in accordance with the release instruction provided during deposition, Figure 16. Release instruction options are summarised in the table below. (Note that the physical release of large entries is not instantaneous. Synchronisation with mirror sites may lead to additional delays before an entry is shown on such sites.)
| Release instruction | Description | |
| REL | As soon as the annotation procedure is complete and the entry has been approved by the depositor, the release procedure will be initiated | |
| EMDBPUB | Release after the associated EMDB entry has been released. If one year after the deposition date the associated EMDB entry has not been released, the EMPIAR entry will be deleted and never be publicly released. (Later release will require the data to be deposited anew.) The EMPIAR accession code will not be recycled. A one-time extension of no more than 6 months will be considered if (one of) the owner(s) requests this and provides a reasonable explanation | |
| HPUB | Release after the primary citation for the dataset becomes available. The same procedure as for EMDBPUB will be applied if the publication is not available one year after the deposition date | |
| HPRE | Release after the preprint citation for the dataset becomes available. The same procedure as for EMDBPUB will be applied if the publication is not available one year after the deposition date | |
| HOLD | Release after a specified period, not to exceed one year. This option is only available if there is no related EMDB entry or publication. A one-time extension of no more than 6 months will be considered if (one of) the owner(s) requests this and provides a reasonable explanation | |

Please note that while we have automated checks in place to find out, for example, when the citation is published, these checks might fail to detect one of these events. We therefore recommend that you contact the EMPIAR annotation team to let them know when associated entries or citations are released/published.
6.4 Upload data
You will not be able to proceed to the Upload data until the Deposition overview page has been completed and validated. You are provided with three options — Globus, Aspera via command line and Aspera via web-client. Once the upload has finished, please check your data on "Associate image sets with data page" as described below.
When using the web-client please keep in mind that there is a limitation set by the web-browser and the operating system for the selection dialogue in the web-browser. Usually the most you can select is about 300 files at a time (depends on the length of filenames and paths to files). If you intend to upload more than that in a single go (as opposed to performing multiple click/select operations to upload 300 files at a time) or if your dataset is 400 GB+ in size, we recommend using the Aspera command line client.
Data-transfers commonly proceed at 50 - 200 GB per hour so expect TB+ sized datasets to take days in some cases. If you are using the command-line client or Globus you can do so asynchronously without being logged in to the deposition system. However you need tokens to initiate the transfers which are provided on the Upload data.
6.5 Associate image sets with data page
Due to the fact that we do not prescribe the organization of data being uploaded, the purpose of this page is to allow the depositor to identify and describe the datasets present in the uploaded data. As an example one could have three datasets — raw multi-frame micrographs, frame-averaged micrographs and particle stacks that have to be associated with the directories "micrographs/multiframe/", "micrographs/singleframe/" and "particles/" respectively. As data upload may proceed asynchronously, you may proceed to this page even though the upload has not completed.
6.5.1 Checking the uploaded data
- The "Refresh directory structure" button will re-build a logical representation of the directory tree structure and determine the size of the upload, Figure 17.
- It will also check for zero sized files and provide warnings if any are found. These are all good initial checks to see if the upload has completed and has been successful.
- A more detailed check can be done by comparing the md5 sums for all the uploaded files with the md5 sums of the files on your local disk. In order to make this check possible we provide a json file and a Python script that can be downloaded and run by you to check that files match. More detailed instructions can be seen by pressing the "Check the uploaded data in EMPIAR" on the image set association page.
- Also the same button gives you an option to download the list of the uploaded files.
- We recommend that you run all these checks to make sure that the data has been uploaded correctly.

6.5.2 Workflow
If you used software that recorded the workflow, for example, Scipion, then you would be able to provide a great way to reproduce previous processing steps and is particularly useful to repeat steps for similar samples or to share knowledge between users.
6.5.3 Associating datasets
- You need to define at least one dataset.
- Press "Set directory" butten, then use the directory tree browser and select the directory corresponding to the dataset, Figure 18. Click on the directory — this will automatically populate the corresponding field in the form.
- Fill out the form fields describing the dataset. Please note that a descriptive name is useful especially when the deposition consists of several datasets. The "Details" section is also useful to describe auxiliary data and how it may be related to the image data.
- You can fill in some of the fields automatically by clicking on one of the image set files, and, if it is readable by IMOD or BSOFT, you will see its header displayed in a popup. There you can click a button to populate all possible fields in the corresponding form, Figure 19.
- To add another dataset, please press "Add more" button at the bottom of the page, Figure 20.



7 Helpdesk

There are three help options available from the left menu, Figure 21. This manual can be accessed from "Deposition manual". The "Helpdesk" link in the left menu can be used to pose a question or review previous communications with the annotation staff. To pose questions specifically about a deposition that is being edited, we recommend that you use the "Deposition help" button. The help desk system allows you to add attachments to your communications. If you have trouble with registering an EMPIAR account or using the helpdesk system, please send us e-mail.
8 Invite reviewers

You may be requested by editors or referees to provide an access to your data before the publication. To facilitate this we provide the owner of the entry with an option to generate credentials for an anonymous user that can be used to log into the EMPIAR deposition system to review your metadata, download and check your data.
