Database of Genomic Variants archive
Phasing out support for the Database of Genomic Variants archive (DGVa).
The submission, archiving, and presentation of structural variation services offered by the DGVa is transitioning to the European Variation Archive (EVA). All of the data shown in the DGVa website is already searchable and browsable from the EVA Study Browser.
Submission of structural variation data to EVA is done using the VCF format. The VCF specification allows representing multiple types of structural variants such as insertions, deletions, duplications and copy-number variants. Other features such as symbolic alleles, breakends, confidence intervals etc., support more complex events, such as translocations at an imprecise position.
We expect to cease accepting direct submissions to DGVa at the end of 2019, in the meantime we
recommend submitters make SV submissions to the EVA. If there are specific difficulties with
preparing SV submissions in VCF format, please contact the EVA helpdesk.
The Database of Genomic Variants archive (DGVa) is a repository that provides archiving, accessioning and distribution of publicly available genomic structural variants, in all species.
In recent years there have been unprecedented advances in the technologies that characterise genomic variation, and it is well known that variation at the single nucleotide level is abundant across the genomes of all species. However, it is becoming clear that genomic structural variation - this is variation ranging from tens to millions of base pairs in size and includes insertions, deletions, inversions, translocations and locus copy number changes - accounts for more of the individual differences at the base pair level in humans and is likely to play a major role in disease. Two other areas of research that are becoming increasingly important in this field are discovering how genomic structural variation affects an individual's characteristics, and understanding the role it has played in the evolution of species. The DGVa catalogues, stores and freely disseminates this important class of variation in any species, providing a valuable resource to a large community of researchers.
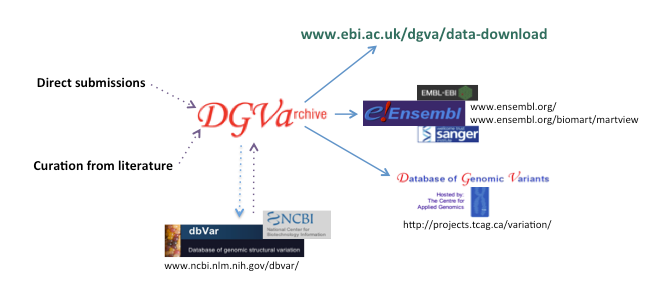
Figure 1. DGVa is a central repository that receives data from, and distributes data to, a number of resources
The DGVa accepts direct submissions from researchers and performs manual curation from the literature. The DGVa also exchanges data on a regular basis with dbVar (a peer archive hosted by NCBI in the USA). Data can be retrieved from DGVa's data download page . Data can be viewed in a richly annotated genomic context using the Ensembl genome browser, or selectively mined and downloaded using Ensembl BioMart. The DGVa also supplies data to DGV (Database of Genomic Variants, hosted by The Centre for Applied Genomics in Canada), where additional annotation and interpretation is performed.
The archive data model and accessioned objects
Stable identifiers (accession numbers) are provided for the STUDY, the genomic region in which the variation occurs (VARIANT REGION) and the particular variant observed in a individual sample (VARIANT CALL). The archive also collects and stores important information relating to those objects, such as sample details, experimental procedures and assertion methods (shown in blue in Figure 2 below.)
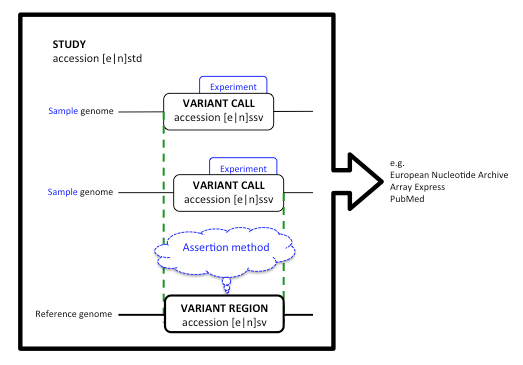
Figure 2. The data model that links accessioned objects
The three types of accessioned objects are prefixed with e if processed by DGVa, n if processed by dbVar. Variation in individual sample genomes is aggregated to a variant region with respect to a reference genome, by procedures described in the Assertion method. Genomic positions of variant calls (shown in green) do not necessarily overlap completely. Discovery and validation procedures are described in the Experiment attribute for each call. The study is the container for all information relating to the body of work and points to any external resources that provide access to raw data (such as the European Nucleotide Archive or Array Express) or to publications describing the study and data (such as PubMed .)
