Exploring phenotypes associated with a variant
To learn more about how rs334 is linked to phenotypes you can search for it in the GWAS Catalog. A search for “rs334” returns the entry for that variant, containing all relevant data in the GWAS Catalog, such as studies, associations and traits associated with rs334 (Figure 8).
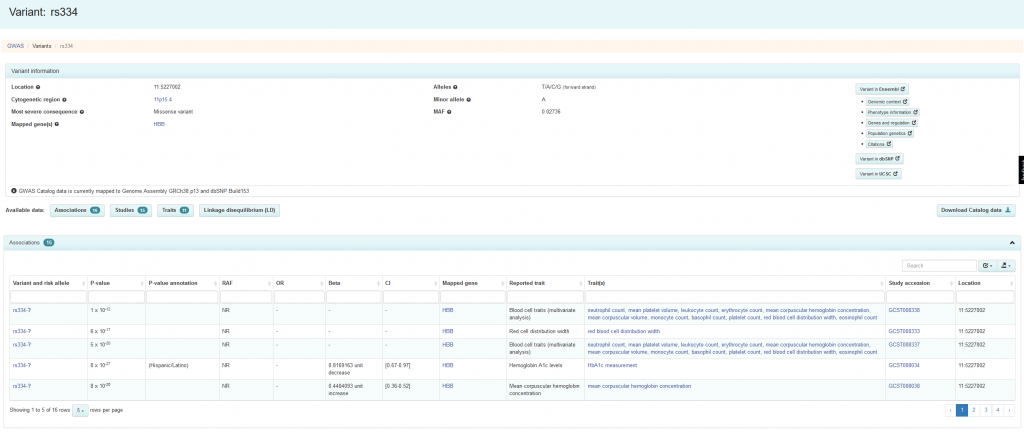
In the GWAS Catalog, rs334 is reported to be associated with urinary albumin-to-creatinine ratio and severe malaria. Summary information is provided for each association, including p-value, effect size (odds ratios or -coefficient), risk-allele, location and mapped gene. The Catalog includes information describing the GWAS in which this association was identified, including links to the publication, study design and ontology terms to describe the phenotype and allow integration with data from other resources. Since GWAS are generally used to study complex, rather than simple Mendelian inheritance, we do not see sickle cell anaemia listed here.
|
|