- Course overview
- Search within this course
- Network analysis in biology
- Introduction to graph theory
- Types of biological networks
- The sources of data underlying biological networks
- Building and analysing PPINs
- Summary
- Quiz: Check your learning
- Your feedback
- Learn more
- References
Properties of PPINs: scale-free networks
Protein-protein interaction networks are scale-free networks (Figure 18A). The majority of nodes (proteins) in scale-free networks have only a few connections to other nodes, whereas some nodes (hubs) are connected to many other nodes in the network.
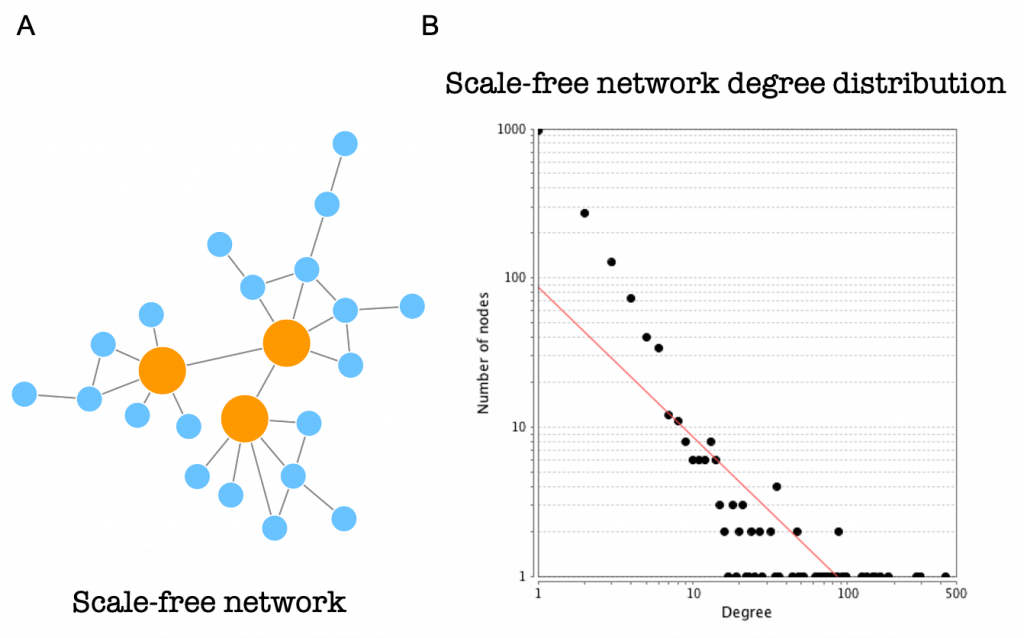
The number of connections each node has is called its degree. If we represent the degree distribution of a scale-free network in a logarithmic scale, we can see how it fits with a line (they fit a power-law), having a small number of nodes with high degree (the hubs) and a large number of nodes with a low degree (Figure 18B).
Scale-free networks can be built following the preferential attachment model, also known as the ‘rich get richer’ principle. This principle simply states that scale-free networks can be built by adding edges that are preferentially attached to those nodes with a highest degree (5). This building principle provides a self-organising mechanism for the generation and expansion of this type of network.
The scale-free nature of protein-protein interaction networks gives them a number of important features:
- Stability
- If failures occur at random, and the vast majority of proteins are those with a small degree of connectivity, the likelihood that a hub would be affected is small
- If a hub-failure occurs, the network will generally not lose its connectedness, due to the remaining hubs
- Invariant to changes of scale
- No matter how many nodes or edges the network has, its properties remain stable
- The presence of hubs is what allows for the small-world effect to be present regardless of the size of the network
- Vulnerable to targeted attack
- If we lose a few major hubs from the network, the network is turned into a set of rather isolated graphs
- Hubs are enriched with essential/lethal genes. For example, many cancer-linked proteins are hub proteins (e.g. the tumour suppressor protein p53)